June 27, 2008
Dairying in Neolithic Central Europe
Direct evidence for the existence of dairying farms in prehistoric Central Europe (4th millennium BC).
Spangenberg JE, Matuschik I, Jacomet S, Schibler J.
The molecular and isotopic chemistry of organic residues from archaeological potsherds was used to obtain further insight into the dietary trends and economies at the Constance lake-shore Neolithic settlements. The archaeological organic residues from the Early Late Neolithic (3922-3902 BC) site Hornstaad-Hornle IA/Germany are, at present, the oldest archaeological samples analysed at the Institute of Mineralogy and Geochemistry of the University of Lausanne. The approach includes (13)C/(12)C and (15)N/(14)N ratios of the bulk organic residues, fatty acids distribution and (13)C/(12)C ratios of individual fatty acids. The results are compared with those obtained from the over 500 years younger Neolithic (3384-3370 BC) settlement of Arbon Bleiche 3/Switzerland and with samples of modern vegetable oils and fat of animals that have been fed exclusively on C(3) forage grasses. The overall fatty acid composition (C(9) to C(24) range, maximizing at C(14) and C(16)), the bulk (13)C/(12)C and (15)N/(14)N ratios (delta(13)C, delta(15)N) and the (13)C/(12)C ratios of palmitic (C(16:0)), stearic (C(18:0)) and oleic acids (C(18:1)) of the organic residues indicate that most of the studied samples (25 from 47 samples and 5 from 41 in the delta(13)C(18:0) vs. delta(13)C(16:0) and delta(13)C(18:0) vs. delta(13)C(18:1) diagrams, respectively) from Hornstaad-Hornle IA and Arbon Bleiche 3 sherds contain fat residues of pre-industrial ruminant milk, and young suckling calf/lamb adipose. These data provide direct proof of milk and meat (mainly from young suckling calves) consumption and farming practices for a sustainable dairying in Neolithic villages in central Europe around 4000 BC
Link
June 26, 2008
Faces close to group prototypes are attractive
Locating attractiveness in the face space: faces are more attractive when closer to their group prototype.
Potter T, Corneille O.
Face attractiveness relates positively to the mathematical averageness of a face, but how close attractive faces of varying groups are to their own and to other-group prototypes in the face space remains unclear. In two studies, we modeled the locations of attractive and unattractive Caucasian, Asian, and African faces in participants' face space using multidimensional scaling analysis. In all three sets of faces, facial attractiveness significantly increased with the absolute proximity of a face to its group prototype. In the case of Caucasian and African faces (Study 1), facial attractiveness also tended to increase with the absolute proximity of a face to the other-group prototype. However, this association was at best marginal, and it became clearly nonsignificant when distance to the own-group prototype was controlled for. Thus, the present research provides original evidence that average features of faces contribute to increasing their attractiveness, but only when these features are average to the group to which a face belongs. The present research also offers further support to face space models of people's mental representations of faces.
Link
June 25, 2008
8,700 year old clock gene selected in non-Africans
Genetic diversity patterns at the human clock gene period 2 are suggestive of population-specific positive selection
Fulvio Cruciani et al.
Abstract
Period 2 (PER2) is a key component of the mammalian circadian clock machinery. In humans, genetic variation of clock genes or chronic disturbance of circadian rhythmicity has been implied in the onset of several phenotypes, ranging from periodic insomnias to advanced or delayed sleep phases, to more severe disorders. Peculiar geographic diversity patterns in circadian genes might represent an adaptive response to different light/dark cycles or environmental changes to which different human populations are exposed. To investigate the degree and nature of PER2 gene variation in human populations of different geographic origin, and its possible correlation with different latitudes, we sequenced a 7.7 kb portion of the gene in 20 individuals worldwide. In total, 25 variable sites were identified. The geographic distribution of haplotypes defined by five polymorphic sites was analyzed in 499 individuals from 11 populations from four continents. No evidence for latitude-driven selective effects on PER2 genetic variability was found. However, a high and significant difference in the geographic distribution of PER2 polymorphisms was observed between Africans and non-Africans, suggesting a history of geographically restricted natural selection at this locus. In support of this notion, we found several signals for selection in the sequences. The putative selected haplotype showed a recent coalescent age (8.7 Kyr), and an unusually high frequency in non-African populations. Overall, these findings indicate that a human clock-relevant gene, PER2, might have been influenced by positive selection, and offer preliminary insights into the evolution of this functional class of genes.
Link
Admixture estimate Vs. Population-of-origin estimate
The classification problem assumes that each individual has a single origin, e.g., "NW European" or "SE European" and attempts to guess this origin.
For example, EURO-DNA-CALC 1.02 is a classifier, and tries to guess an individual's origin out of the set {"NW EUropean", "SE European", "Ashkenazi Jewish"}.
The admixture estimate problem assumes that each individual is a mixture of groups, and attempts to guess the relative strength of the components in the mixture.
EURO-DNA-CALC 1.1.1 is an admixture estimator, and tries to guess an individual's admixture from the groups "NW EUropean", "SE European", "Ashkenazi Jewish".
In many of the testees that have communicated their results to me, the classification provided by 1.0.2 is equal to the most important component identified by 1.1.1. But this does not need to be the case.
To understand why, let's consider some real-life examples.
The Proto-Uralics
Using skull measurements, scientists have identified traits that seem to have belonged to a Proto-Uralic human group. Presently, individuals of Northern Eurasia, especially Uralic speakers are a mix of Western Eurasian (Caucasoid) and Eastern Eurasian (Mongoloid) elements, but they also possess a small and varying component of this old Proto-Uralic group.
So, an individual could be guessed as a Uralic speaker with some success, and yet his Uralic admixture would be generally low.
African Americans
It is well-known that African Americans have European admixture which varies a lot individually. Indeed, some African Americans are more European genetically than they are African. On the other hand, European Americans generally tend to have insignificant African admixture. This assymetry is due to the fact that individuals of partial African ancestry were reckoned to be "Black", no matter what the exact percentage of their African ancestry.
Now, a classifier that would distinguish between Whites and African Americans would have no trouble distinguishing between the two groups easily. But this classification would not need to be concordant with their admixture estimate:
A person of 25% African ancestry is extremely more likely to be (in the US) an African American than a White. And yet, the high classification probability that he is African American is discordant with their most important component, i.e., European.
The "Brown" Cluster
In the recent paper on human genetic variation using 650K SNPs, a "Brown" cluster was identified. Middle Eastern populations seemed to have a high fraction of their ancestry from this cluster:

If we built a classifier to distinguish between Europeans and non-Europeans, the presence of the "Brown" cluster would give us near certainty that an individual is non-European and yet there are non-European individuals (e.g., the Druze), where the "European" component (average for the group 52%) is greater than the "Brown" one (34%).
June 24, 2008
The eclipse of 1178BC and the return of Odysseus to Ithaca
 Odysseus' return from Trojan War dated:
Odysseus' return from Trojan War dated:In the epic "Odyssey," one of the cornerstones of Western literature, the legendary Greek hero Odysseus returns to his queen Penelope after enduring 10 years of sailing the wine dark sea.
Now scientists have pinned down his return to April 16, 1178 B.C., close to noon local time, according to astronomical references in the epic poem that seem to pinpoint the total eclipse of the sun on the day that Odysseus supposedly returned on.
...
The possible solar eclipse comes up in the 20th book of the "Odyssey," as the suitors begin their final lunch. At this point, the goddess of war Athena "confounds their minds," making the suitors laugh uncontrollably and see their food spattered with blood. The seer Theoclymenus then foresees the death of the suitors, ending by saying, "The sun has been obliterated from the sky, and an unlucky darkness invades the world."
The Greek historian Plutarch suggested the prophecy of Theoclymenus referred to a solar eclipse.
More recently, astronomers Carl Schoch and Paul Neugebauer computed in the 1920s that a total solar eclipse occurred over the Ionian islands — of which Ithaca is one — about noon on April 16, 1178 B.C., and would have coincided roughly a decade before the most often cited estimate for the sack of Troy — about 1190 B.C.
...
Still, a great deal of skepticism remains over whether Theoclymenus refers to this or any eclipse. To shed light on the issue, researchers Marcelo Magnasco and Constantino Baikouzis at Rockefeller University in New York decided to analyze other passages in the "Odyssey" for astronomical references without assuming an eclipse.
...
The scientists first created a rough chronology of events depicted in the "Odyssey." First, 29 days before the supposed eclipse and the massacre of the suitors, three constellations are mentioned as Odysseus sets out from the island of Ogygia, where he has spent seven years as a captive of the beautiful nymph Calypso. Odysseus is told to watch the Pleiades and late-setting Boötes and keep the Great Bear to his left. Next, five days before the supposed eclipse, Odysseus arrives in Ithaca as the Star of Dawn — that is, Venus — rises ahead of the sun.Finally, the night before the eclipse, there is a new moon.
Also, the messenger of the gods, Hermes, is sent west to Ogygia by the king of the gods Zeus to release Odysseus and then immediately returns back east roughly 34 days before the eclipse. The researchers conjecture this trip refers to an apparent turning point of the motion of the planet Mercury. (Mercury was the Roman name for Hermes.)
...
The scientists then searched for potential dates that satisfied all these astronomical references close to the fall of Troy, which has over the centuries been estimated to have occurred between roughly 1250 to 1115 B.C. From these 135 years, they found just one date satisfied all the references — April 16, 1178 B.C., the same date as the proposed eclipse.
...
"That's just one day out of about 50,000 days," Magnasco told LiveScience. "If our findings are correct, it would be pretty spectacularly strange. How could Homer have known about this eclipse, about planetary positions that happened some 100 years before him? If this is all true, it would change the timetable of what we think they knew about astronomy then." Homer, if he really existed, is said to have composed the "Odyssey" sometime near the end of the ninth century B.C.
Proc. Natl. Acad. Sci. USA, 10.1073/pnas.0803317105
Is an eclipse described in the Odyssey?
Constantino Baikouzis, and Marcelo O. Magnasco
Abstract
Plutarch and Heraclitus believed a certain passage in the 20th book of the Odyssey ("Theoclymenus's prophecy") to be a poetic description of a total solar eclipse. In the late 1920s, Schoch and Neugebauer computed that the solar eclipse of 16 April 1178 B.C.E. was total over the Ionian Islands and was the only suitable eclipse in more than a century to agree with classical estimates of the decade-earlier sack of Troy around 1192–1184 B.C.E. However, much skepticism remains about whether the verses refer to this, or any, eclipse. To contribute to the issue independently of the disputed eclipse reference, we analyze other astronomical references in the Epic, without assuming the existence of an eclipse, and search for dates matching the astronomical phenomena we believe they describe. We use three overt astronomical references in the epic: to Boötes and the Pleiades, Venus, and the New Moon; we supplement them with a conjectural identification of Hermes's trip to Ogygia as relating to the motion of planet Mercury. Performing an exhaustive search of all possible dates in the span 1250–1115 B.C., we looked to match these phenomena in the order and manner that the text describes. In that period, a single date closely matches our references: 16 April 1178 B.C.E. We speculate that these references, plus the disputed eclipse reference, may refer to that specific eclipse.
Link
June 23, 2008
The Tuscans (Validation of EURO-DNA-CALC)
My first test was on the 8 Tuscan individuals of the Stanford dataset. Here are their individual admixture estimates (NW Euro, SE Euro, Ashkenazi Jewish)
| 9 | 72 | 19 |
| 24 | 70 | 6 |
| 25 | 75 | 0 |
| 3 | 97 | 0 |
| 0 | 100 | 0 |
| 0 | 100 | 0 |
| 16 | 84 | 0 |
| 13 | 87 | 0 |
The average results for the Tuscans are in the pie graph:
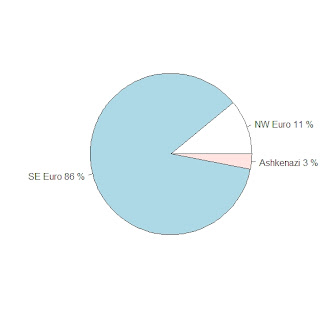
These results seem reasonable; at least we can conclude that EURO-DNA-CALC correctly guessed the Tuscans as Southeastern European individuals.
June 21, 2008
EURO-DNA-CALC 1.1 released
A new version of the EURO-DNA-CALC has been released. The previous version (1.0) was described here.
Version 1.1 also uses deCODEme/23andMe data for an individual but outputs a maximum likelihood estimate of his admixture proportions from the three groups NW European, SE European, and Ashkenazi Jewish.
More comments on this dna-forums thread (registration required).
Below are the admixture estimates for Greg and Lily Mendel of 23andMe. An examination of the confidence intervals output by the program suggests that non-NW European admixture in Greg is a strong possibility, while Lily could conceivably have no non-NW European admixture. On the other hand, Lily's 10% Ashkenazi and only 1% SE European admixture imbalance is interesting. As with all genetic tools, the results should be interpreted in the light of other information about a person's ethnic origin and genealogy.


June 19, 2008
IGF1, ESR2, and CYP17 genes and adult height in Caucasians
Comprehensive association analyses of IGF1, ESR2, and CYP17 genes with adult height in Caucasians
Tie-Lin Yang et al.
Human adult height is closely related to body growth that is regulated by multiple cytokines or hormones like growth hormone (GH) and estrogen. Our study focused on three potential candidate genes to human height, namely IGF1 (insulin-like growth factor 1), ESR2, and CYP17. We genotyped 43 single nucleotide polymorphisms (SNPs) and tested their associations in 1873 subjects from 405 nuclear families, using both the family-based quantitative transmission disequilibrium test (QTDT) and population-based ANOVA methods. Both analyses consistently detected that two novel SNPs of IGF1, rs5742694 and rs2033178, were significantly associated with human height, with the P-values of 0.0097 and 0.0057 in QTDT analyses, 0.0002/0.004 (sample 1/sample 2) and 8.46 times 10- 5/1.92 times 10- 5 in ANOVA analyses. For ESR2, significant associations were only detected in women (rs1256061: QTDT P=0.002, ANOVA P=0.002/0.012; rs17766755: QTDT P=0.019, ANOVA P=0.023/0.006; rs1256044: QTDT P=0.022, ANOVA P=0.002/0.034). Haplotype analyses corroborated our single-SNP results. However, no association was detected between CYP17 and human height. In conclusion, we identified the important effects of IGF1 and ESR2 on adult height variation in Caucasians, and first suggested the potential sex-specific effect of ESR2 on height variation in Caucasian women. It will be valuable for other independent studies to replicate and confirm these findings.
Link
June 18, 2008
Scanning the human genome at kilobase resolution
Genome Res. 2008 May;18(5):751-62. Epub 2008 Feb 21.
Scanning the human genome at kilobase resolution.
Chen J, Kim YC, Jung YC, Xuan Z, Dworkin G, Zhang Y, Zhang MQ, Wang SM.
Normal genome variation and pathogenic genome alteration frequently affect small regions in the genome. Identifying those genomic changes remains a technical challenge. We report here the development of the DGS (Ditag Genome Scanning) technique for high-resolution analysis of genome structure. The basic features of DGS include (1) use of high-frequent restriction enzymes to fractionate the genome into small fragments; (2) collection of two tags from two ends of a given DNA fragment to form a ditag to represent the fragment; (3) application of the 454 sequencing system to reach a comprehensive ditag sequence collection; (4) determination of the genome origin of ditags by mapping to reference ditags from known genome sequences; (5) use of ditag sequences directly as the sense and antisense PCR primers to amplify the original DNA fragment. To study the relationship between ditags and genome structure, we performed a computational study by using the human genome reference sequences as a model, and analyzed the ditags experimentally collected from the well-characterized normal human DNA GM15510 and the leukemic human DNA of Kasumi-1 cells. Our studies show that DGS provides a kilobase resolution for studying genome structure with high specificity and high genome coverage. DGS can be applied to validate genome assembly, to compare genome similarity and variation in normal populations, and to identify genomic abnormality including insertion, inversion, deletion, translocation, and amplification in pathological genomes such as cancer genomes.
Link
Women with high-pitched voices are more attractive
The role of femininity and averageness of voice pitch in aesthetic judgments of women's voices.
Feinberg DR, DeBruine LM, Jones BC, Perrett DI.
Although averageness is preferred in auditory stimuli (eg music) and non-face objects (eg wristwatches), exaggerated feminine characteristics are preferred to averageness in female faces. To establish whether or not men prefer femininity in female voices to average characteristics, we conducted a correlational study (study 1) to assess the relationship between voice pitch and attractiveness ratings. We found a positive linear relationship between voice pitch and attractiveness ratings. In study 2 we manipulated pitch in women's voices with low (lower than average), average, and high (higher than average) starting pitches and gauged men's preferences. Men preferred women's voices with raised pitch for all levels of starting pitch. These findings suggest that men prefer high voice pitch to average voice pitch in women's voices.
Link
June 16, 2008
EURO-DNA-CALC 1.0 released
I have created a version of my classifier for 23andMe/deCODEme genotype data that I talked about before. It calculates the probability that an individual is "Northwestern European", "Southeastern European", or "Ashkenazi Jewish".
You can download it here.
If you use it, feel free to leave a comment or send me an e-mail.
Comments from some users who tried it can also be found in this dna-forums thread (registration required).
(clarification, June 17): This is not an admixture test, but a "guess his/her origin out of these three groups" test, a classifier. Here is a way to look at it: the classifier compares these three events:
How likely is this genotype to be NW European?
How likely is this genotype to be SE European?
How likely is this genotype to be Ashkenazi Jewish?
An admixture test compares the likelihood of all possible events:
How likely is this genotype to be (x% NWE, y% SEE, z% AJ), for all x,y,z>0 such as x+y+z=100
Admixture analysis is a harder task than classification, because it looks at a much bigger realm of possibilities (all x,y,z combinations). That is why people have been getting generally accurate assessment of their main ancestry component from various testing services over the years, but sprinkled with widely varying and often puzzling estimates of minor admixture.
June 15, 2008
Haplogroup prediction from Y-STR data
PLoS Comput Biol. 2008 Jun 13;4(6):e1000093.
Machine-learning approaches for classifying haplogroup from Y chromosome STR data.
Schlecht J, Kaplan ME, Barnard K, Karafet T, Hammer MF, Merchant NC.
Genetic variation on the non-recombining portion of the Y chromosome contains information about the ancestry of male lineages. Because of their low rate of mutation, single nucleotide polymorphisms (SNPs) are the markers of choice for unambiguously classifying Y chromosomes into related sets of lineages known as haplogroups, which tend to show geographic structure in many parts of the world. However, performing the large number of SNP genotyping tests needed to properly infer haplogroup status is expensive and time consuming. A novel alternative for assigning a sampled Y chromosome to a haplogroup is presented here. We show that by applying modern machine-learning algorithms we can infer with high accuracy the proper Y chromosome haplogroup of a sample by scoring a relatively small number of Y-linked short tandem repeats (STRs). Learning is based on a diverse ground-truth data set comprising pairs of SNP test results (haplogroup) and corresponding STR scores. We apply several independent machine-learning methods in tandem to learn formal classification functions. The result is an integrated high-throughput analysis system that automatically classifies large numbers of samples into haplogroups in a cost-effective and accurate manner.
Link
June 13, 2008
mtDNA of Tarim mummies




The National Geographic documentary on the Tarim mummies reports on recent mtDNA work on the Tarim mummies. The program doesn't really reveal anything new to anyone familiar with the story of these mummies, but there are some nice segments of some of them as they would have been during their lifetime. At some point, the camera shows what appear to be haplogroup assignments, although I wouldn't vouch as to what these actually mean. or to who exactly they belong. What they do say is that they found markers from "Europe, West Eurasia, Siberia, Tibet, Mongolia, even India". They also mention that the "Beauty of Loulan", the "Boy" have "unexpected marks of East Asian ancestry", and "Cherchen Man" also carries "a surprising East Asian lineage" and that the "Shaman" has a "lineage frequently seen in the Himalayas and India".
Ancient mtDNA from Sampula population in Xinjiang
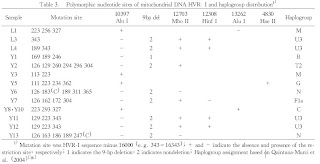 From the paper:
From the paper:Physical anthropology of Shao et al. revealed that the ancient human bones from Sampula exhibited primarily Mongoloid characteristics with certain European features, but Han et al. believed that Sampula populations are mainly of European character and actually are close to that of the Eastern Mediterranean type.
...
In conclusion, the analysis of mtDNA haplogroup distribution showed that the ancient Sampula was a complex population of European and Asian, corresponding to the physical anthopology result of Shao et al.
Progress in Natural Science, Volume 17, Issue 8 August 2007 , pages 927 - 933
Mitochondrial DNA analysis of ancient Sampula population in Xinjiang
Chengzhi Xie et al.
Abstract
The archaeological site fo Sampula cemetery was located about 14 km to the southwest of the Luo County in Xinjiang Khotan, China, belonging to the ancient Yutian kingdom. 14C analysis showed that this cemetery was used from 217 B.C. to 283 A. D. Ancient DNA was analysed by 364 bp of the mitochondrial DNA hypervariable region 1 (mtDNA HVR-1), and by six restriction fragment length polymorphism (RFLP) sites of mtDNA coding region. We successfully extracted and sequenced intact stretches of maternally inherited mtDNA from 13 out of 16 ancient Sampula samples. The analysis of mtDNA haplogroup distribution showed that the ancient Sampula was a complex population with both European and Asian Characteristics. Median joining network of U3 sub-haplogroup and multi-dimensional scaling analysis all showed that the ancient Sampula had maternal relationship with Ossetian and Iranian.
Link
June 12, 2008
Classifier for 23andMe/deCODEme genotype data
I have built a simple classifier tool based on the panel of 300 markers of Price et al. (2008), which uses the frequency data supplied in this paper to assess the probability that an individual belongs to the "Northwest European", "Southeast European", or "Ashkenazi Jewish" categories.
The input is genotype values for a number of markers (e.g., the 169/192 markers in common between the deCODEme and 23andMe results) for an individual, and the output is a set of three probabilities for belonging to any of the three groups, summing up to 1.
Using the Greg and Lilly Mendel data that you can download from 23andMe, I came up with the following probabilities (NWE,SEE,AJ):
Greg: 0.89, 0.11, 0
Lilly: 1, 0, 0
(corrected June 15)
23andMe lists the similarity of these individuals to "Northern Europeans","Southern Europeans", and "Near Easterners" as:
Greg: 67.84, 67.74, 67.15
Lilly: 67.85, 67.72, 67.11
So, at least for these two individuals the results of my calculator appear to be analogous to those reported by 23andMe, with Lilly seeming more "Northern" than her husband.
PS: Unfortunately my calculator cannot be released at present, as it's not a standalone program but rather relies on a bunch of different tools with minimum development.
June 11, 2008
deCODEme, 23andMe SNPs and published Caucasoid substructure studies
Both services seem to use the HGDP populations for the more specific (subracial) similarity assessment. In the last couple of years, there have been a few studies that looked at intra-European or intra-Caucasoid genomic variation, so it might be possible to devise a test using the published results and the SNPs tested by these companies.
deCODEme uses the Illumina 1M BeadChip, while 23andMe uses the Illumina HumanHap550+ BeadChip with an additional custom set of markers. The deCODEme chip measures 1,072,820 SNPs, while 23andME (according to the "Greg Mendel" data you can download from their website) measures 571,754 SNPs.
Price et al. (2008) have identified a set of 300 ancestry informative markers including that distinguishes between NW/SE Europeans and SE Europeans/Ashkenazi Jews. The deCODEme set tests for 192 of these markers, whereas 23andME tests for 169 of them.
Tian et al. (2008) have identified 1,441 European substructure ancestry informative markers (rtf) (ESAIMs). deCODEme tests 1,412 of them, while 23andME tests 1,424 of them. As far as I can tell, the original study did not publish either frequency data or individual genotypes for these markers, so using them to infer ancestry may not be possible. (Let me know if this data exists and I missed it).
(added Jun 12) Bauchet et al. (2007) have identified a panel of 1,200 markers for European population substructure and report frequency data for Southeastern and Northern Europeans (xls). deCODEme tests for 508 of them, and 23andMe tests for 438 of them.
Seldin et al. (2006) have provided frequency data (pdf) in several populations for 5,735 SNPs. markers. deCODEme tests for 3,502 of these, while 23andMe tests for 2,242 of them.
One could also use the 650K SNPs from Stanford. There are 660,755 non-mitochondrial SNPs in the freely available data, all of which are tested by deCODEme; 23andMe which partially funded the study (Li et al. (2008)) tests 549,118 of them.
In conclusion, it seems possible to make your own test using commercially available SNPs and freely available data other than the HGDP populations.
June 10, 2008
Craniometry of prehistoric populations of the south-central Andes
The genetic divergence of prehistoric populations of the south-central Andes as established by means of craniometric traits
Héctor H. Varela et al.
Abstract
The peopling of the south-central Andean region can be determined by exploring a combination of cultural, economic, and biological factors that influence the structure of populations and determine particular dispersals of gene frequencies. Quantitative characters from 1,586 adult crania of both sexes from northern Chile, northwestern Argentina, and the Cochabamba valleys in Bolivia were analyzed employing multivariate statistical analyses. Biological distances, representing phenotypic variation between these regions and their subregions, were studied within a population genetics framework. An analysis of Mahalanobis D2 distances establishes two principle directions of interaction: the first between the Cochabamba valleys and northern Chile, and the second between the Cochabamba region and northwestern Argentina. The Chile and Argentina regions are shown to be less related to each other than each is to the Bolivian region. A higher mean genetic divergence is found for the entire region (FST = 0.195); with northwestern Argentina having the highest spatial isolation (FST = 0.143) and northern Chile the lowest (FST = 0.061). These results allow us to propose a populating model based on the dispersion of several lines from a common ancestral population similar to those who inhabited the Cochabamba valleys. These lines differentiated themselves in time and space according to the effective size and the rate of gene flow, eventually producing the human groups which inhabited the valleys of northern Chile and northwestern Argentina.
Link
June 09, 2008
The human settlement in Eurasia: The mountainous Central Asia and the Sub-Himalayan piedmonts
Face à une telle dispersion dans le temps et l’espace, la présence du torus angularis sur des crânes néolithiques indiens peut difficilement s’expliquer autrement que par une parenté génétique avec des Homo erectus indiens. Une origine asiatique de l’Homo sapiens est tout à fait vraisemblable. Cette étude devrait se compléter de la comparaison des populations harappéennes entreposées à Calcutta (civilisation chalcolithique de l’Indus). Il serait intéressant de vérifier si le torus angularis est également présent sur les crânes néolithiques de Merghar (Pakistan) et sur le crâne de Pahiyangala découvert dans une grotte au Sri Lanka et daté de 37 000 ans.
L'Anthropologie doi:10.1016/j.anthro.2008.04.008
Le peuplement humain en Eurasie : l’Asie centrale montagneuse et les piémonts sous-himalayens du Plio-Pléistocène à l’Holocène, origines, évolution humaine et migrations
Anne Dambricourt Malassé
Abstract
During the years 1996 and 1997, a team of the Laboratory of Prehistory, National Museum of Natural History, Paris, and of the Departments of Archaeology, Karachi and Peshawar University, Pakistan, leads the first prehistoric field investigation in the District of Chitral, Hindu Kush, close to the Wakhan Corridor (the Amu Daria course in the Pamir). Problematics are the origins and the becoming of the Epipaleolithic/Neolithic hunters-gatherers known in the Pamir Plateau and the Gissar Range, the lithics tradition of which share common roots with the Sub-Himalayan Soanian tradition (Mode 1). A second field investigation has been conduced in the North West India, where Soan developed from Early Pleistocene, in the Frontal Range of the Siwaliks and Himachal Pradesh during the years 2003, 2005 and 2006 in cooperation with the Department of Archaeology and Museums of Punjab, India. New discoveries in both countries support new hypothesis for the understanding of human evolution in Asia and Homo sapiens origins.
LinkJune 08, 2008
Stature increase in China
Secular changes in stature and body mass index for Chinese youth in sixteen major cities, 1950s-2005
Cheng-Ye Ji et al.
Abstract
Evidence shows a secular trend in physical growth in China in recent years. We analyze the secular trend of stature and body mass index (BMI) for the period 1950s-2005 to provide biological evidence for policy-makers to identify measures for improving Chinese children's health. Data come from the historical records in 1950s and the successive cycles of the Chinese National Survey on Student's Constitution and Health. Subjects were 7- to 18-year-old youth from 16 cities. Sex-age differences in mean stature and BMI values between the surveys were analyzed, and the increments per decade were compared. An overall positive secular trend was found in 1950s-2005. Mean stature of the 18-year olds increased from 166.6 to 173.4 cm for males and from 155.8 to 161.2 cm for females, yielding rates of 1.3 and 1.1 cm/decade; the overall increments of BMI values were 2.6 for males and 1.8 for females, yielding rates of 0.8 and 0.6/decade, respectively. The most significant changes occurred during puberty. The overall positive secular trend is closely associated with the socioeconomic progress and the improvement of livelihood. Strong evidence suggests that in China this trend will be continued for many years. Further studies are needed to explore how to ensure healthy changes for poorer rural youth. Effective preventive strategies and measures should be taken to prevent the progressive increase in the prevalence of childhood obesity accompanying this trend.
Link
June 07, 2008
Einkorn wheat domestication in Karacadağ
In conclusion, Fig. 1C shows a 2-fold scenario: Einkorn domestication near the Karacadağ and T. urartu domestication along the middle Euphrates. These two domestication events would have met over time and mixed plant remains would occur at the respective sites, such as Dja’de, as the preliminary data of Willcox (2005) might imply. The crossing barrier between these two species could explain why the integrities of the species are maintained. And it also explains why an extinct (Fuller 2007) domesticated 2-grained “Einkorn” wheat (which we assume to be T. urartu) might have followed the spread of agriculture, for example into the Balkans (Kroll 1992) and Germany (Kreuz and Boenke 2002), and would have left no DNA trace in modern Einkorn. Also interesting is the human dimension: these two domestication events meet close to Göbekli Tepe, the impressive site built by hunter-gatherers on the verge of becoming farmers (Schmidt 2007a, 2007b; Curry 2008). One might also put it the other way around: domestication spread north and south from Göbekli Tepe, making the ceremonial meetings at Göbekli Tepe the “spiritual” source of these two domestications. Also, emmer wheat (T. dicoccum) has its domestication site nearby (Luo et al. 2007).Genome. 2008 Jun;51(6):444-451.
Reassessing domestication events in the Near East: Einkorn and Triticum urartu.
Heun M, Haldorsen S, Vollan K.
To reassess domestication events in the Near East, accessions of Triticum urartu from a well-described sampling were combined with a representative sample covering the Karacadağ Einkorn wheat domestication. The observed DNA separation between the two wheat species accounts for the main differentiation, but geographic variation within T. urartu is evident and so is the domestication scenario among wild, feral, and domesticated Einkorn. In contrast to the clear DNA differences, it is difficult to separate living T. urartu from wild Einkorn based on morphology. With archaeobotanical material a distinction of carbonized remains of these two wheats is considered to be impossible. We reviewed the differences concerning morphology and maturity and combined these observations with information about archaeological sites in the Near East. In conclusion, the excavation sites in the middle Euphrates may contain T. urartu rather than Einkorn wheat and T. urartu may underlie the reported occurrence of the extinct 2-grained domesticated "Einkorn" wheat. The first Einkorn wheat domestication sensu stricto seems to have happened around the Karacadağ, as reported earlier. The human dimension shown by the excavation of Göbekli Tepe can explain why domesticated phenotypes might have spread quickly.
Link
June 06, 2008
ASPM, MCPH1, CDK5RAP and BRCA1 and general cognition, reading or language
- Microcephalin and ASPM do not Account for Brain Size Variability
- ASPM and the alphabet
- Has ASPM been the target of recent selection? (*)
- ASPM and Microcephalin don't make people smarter
Intelligence doi:10.1016/j.intell.2008.04.001
Recently-derived variants of brain-size genes ASPM, MCPH1, CDK5RAP and BRCA1 not associated with general cognition, reading or language
Timothy C. Bates et al.
Abstract
Derived changes in genes associated with primary microcephaly (MCPH) have been suggested to be “currently sweeping to fixation” i.e., increasing in frequency in most populations, with the likely outcome that the derived allele will completely displace the ancestral allele over time. Possible causes for this sweep include effects on human reasoning and language. Here we test the hypothesis that these derived alleles are associated with current variation in spoken or written language and related traits. The association of derived alleles of the ASPM, MCPH1, CDK5RAP2 and BRCA1 genes was tested against well-validated measures of dyslexia, specific language impairment, working memory, IQ, and head-size in a family-based association study of over 1776 subjects from 789 families of twins. No evidence for association was found for any gene to any trait. The results strongly did not support the hypothesis that derived alleles in MCPH-related genes are related to the evolution of human language or cognition. Results were compatible with the alternate hypothesis, suggesting that adaptations in these genes associated with a dramatic increase in brain size have long since reached fixation and are now maintained by stabilizing selection.
Link
June 05, 2008
500K SNP study of Oceanian populations
Molecular Biology and Evolution, doi:10.1093/molbev/msn128
Gene Flow and Natural Selection in Oceanic Human Populations, Inferred from Genome-wide SNP Typing
Ryosuke Kimura et al.
It is suggested that the major prehistoric human colonizations of Oceania occurred twice, namely, about 50,000 and 4,000 years ago. The first settlers are considered as ancestors of indigenous people in New Guinea and Australia. The second settlers are Austronesian-speaking people who dispersed by voyaging in the Pacific Ocean. In this study, we performed genome-wide SNP typing on an indigenous Melanesian (Papuan) population, Gidra, and a Polynesian population, Tongans, by using the Affymetrix 500K assay. The SNP data were analyzed together with the data of the HapMap samples provided by Affymetrix. In agreement with previous studies, our phylogenetic analysis indicated that indigenous Melanesians are genetically closer to Asians than to Africans and European Americans. Population structure analyses revealed that the Tongan population is genetically originated from Asians at 70% and indigenous Melanesians at 30%, which thus supports the so-called "Slow train" model. We also applied the SNP data to genome-wide scans for positive selection by examining haplotypic variation, and identified many candidates of locally selected genes. Providing a clue to understand human adaptation to environments, our approach based on evolutionary genetics must contribute to revealing unknown gene functions as well as functional differences between alleles. Conversely, this approach can also shed some light onto the invisible phenotypic differences between populations.
Link
June 04, 2008
Rats and the dating of the earliest colonization of New Zealand
PNAS | June 3, 2008 | vol. 105 | no. 22 | 7676-7680
Dating the late prehistoric dispersal of Polynesians to New Zealand using the commensal Pacific rat
Janet M. Wilmshurst et al.
The pristine island ecosystems of East Polynesia were among the last places on Earth settled by prehistoric people, and their colonization triggered a devastating transformation. Overhunting contributed to widespread faunal extinctions and the decline of marine megafauna, fires destroyed lowland forests, and the introduction of the omnivorous Pacific rat (Rattus exulans) led to a new wave of predation on the biota. East Polynesian islands preserve exceptionally detailed records of the initial prehistoric impacts on highly vulnerable ecosystems, but nearly all such studies are clouded by persistent controversies over the timing of initial human colonization, which has resulted in proposed settlement chronologies varying from {approx}200 B.C. to 1000 A.D. or younger. Such differences underpin radically divergent interpretations of human dispersal from West Polynesia and of ecological and social transformation in East Polynesia and ultimately obfuscate the timing and patterns of this process. Using New Zealand as an example, we provide a reliable approach for accurately dating initial human colonization on Pacific islands by radiocarbon dating the arrival of the Pacific rat. Radiocarbon dates on distinctive rat-gnawed seeds and rat bones show that the Pacific rat was introduced to both main islands of New Zealand {approx}1280 A.D., a millennium later than previously assumed. This matches with the earliest-dated archaeological sites, human-induced faunal extinctions, and deforestation, implying there was no long period of invisibility in either the archaeological or palaeoecological records.
Link
June 03, 2008
Invention of the naval ram
The Earliest Naval Ram
Samuel Mark
Analyses of the Kuyunjik (Kouyunjik) relief and other data suggest Phoenicia probably did not build ships with rams before the Battle of Salamis. A review of Greek literature, iconography, and archaeology suggests the naval ram may have been a Greek invention, appearing at the earliest in the 6th century BC and possibly as late as the 5th century. Its evolution may have led to a shift from laced to pegged mortise-and-tenon joinery in Greek shipbuilding as well as the development of the wineglass-shaped hull and heavier framing. It may also have influenced the development of large-scale bronze-casting in Greece.
Link
June 02, 2008
Strong Reciprocity and the Emergence of Large-Scale Societies
DOI: 10.1177/0048393108315509
Strong Reciprocity and the Emergence of Large-Scale Societies
Benoît Dubreuil
The paper defends the idea that strong reciprocity, although it accounts for the existence of deep cooperation among humans, has difficulty explaining why humans lived for most of their history in band-size groups and why the emergence of larger societies was accompanied by increased social differentiation and political centralization. The paper argues that the costs of incurring an altruistic punishment rise in large groups and that the emergence of large-scale societies depends on the creation of institutions that render control of these costs possible through a social division of sanction.
Link
Young people find oblique eye axis more attractive
Aesthetic Plast Surg. 2008 May 28.
The "Jaguar's Eye" as a New Beauty Trend? Age-Related Effects in Judging the Attractiveness of the Oblique Eye Axis.
BACKGROUND: The eye area plays an important role in the assessment of a person. Although plastic surgery in this area is quite common, only a few studies have evaluated the features that create an attractive eye. This study aimed to determine whether a preference exists for a certain eye axis. METHODS: The stimulus material comprised portrait images of seven women. Two versions of each face were generated that differed only in the position of the eye axis (normal vs rotated). The eye axis was rotated by raising the lateral canthus of the eye about 5 degrees . A total of 250 experimental subjects ranging in age from 15 to 84 years stated their preferential position of the eye axis (horizontal vs oblique). RESULTS: Clear evidence showed that age has an impact on the type of eye axis preferred. Young subjects (age, /=50 years) preferred the eyes more horizontal (p < href="http://www.ncbi.nlm.nih.gov/pubmed/18506510?dopt=Abstract">Link
June 01, 2008
Wise words on Y chromosome phylogeography
How can you determine when and where a lineage originated? And how does the origin and spread of a lineage relate to what we think of as the origin of a population? These are rather contentious issues. According to the simplest way of thinking, current high frequency and high diversity may mark the place of origin of a lineage; but high frequency can also arise by genetic drift, and high diversity by admixture. The time can be calculated in several ways, and a wide range of mutation rates can be used, so molecular dates are much less certain than archaeological ones. In thinking about the second question, we can paraphrase the Italian geneticist Guido Barbujani: imagine that at some time in the future Indian astronauts colonise Mars, and geneticists then type their Y chromosomes. We may well find that their lineages date back to 9,000–20,000 years ago. But we would not be wise to infer that they have been living on Mars for 9,000 years.
