January 31, 2006
Genes for human height
Genetic linkage of human height is confirmed to 9q22 and Xq24
Yao-Zhong Liu et al.
Abstract Human height is an important and heritable trait. Our previous two genome-wide linkage studies using 630 (WG1 study) and an extended sample of 1,816 Caucasians (WG2 study) identified 9q22 [maximum LOD score (MLS)=2.74 in the WG2 study] and preliminarily confirmed Xq24 (two-point LOD score=1.91 in the WG1 study, 2.64 in the WG2 study) linked to height. Here, with a much further extended large sample containing 3,726 Caucasians, we performed a new genome-wide linkage scan and confirmed, in high significance, the two regions’ linkage to height. An MLS of 4.34 was detected on 9q22 and a two-point LOD score of 5.63 was attained for Xq24. In an independent sub-sample (i.e., the subjects not involved in the WG1 and WG2 studies), the two regions also achieved significant empirical P values (0.002 and 0.004, respectively) for “region-wise” linkage confirmation. Importantly, the two regions were replicated on a genotyping platform different from the WG1 and WG2 studies (i.e., a different set of markers and different genotyping instruments). Interestingly, 9q22 harbors the ROR2 gene, which is required for growth plate development, and Xq24 was linked to short stature. With the largest sample from a single population of the same ethnicity in the field of linkage studies for complex traits, our current study, together with two previous ones, provided overwhelming evidence substantiating 9q22 and Xq24 for height variation. In particular, our three consecutive whole genome studies are uniquely valuable as they represent the first practical (rather than simulated) example of how significant increase in sample size may improve linkage detection for human complex traits.
Link
January 30, 2006
Earwax type determined by single letter of DNA
Earwax (cerumen) is a secretory product of ceruminous apocrine glands. Human earwax is a mendelian trait consisting of wet and dry types1. The wet earwax is brownish and sticky, whereas the dry type lacks cerumen. The wet cerumen phenotype is completely dominant to the dry type. The dry type is seen frequently (80–95%) among East Asians2, 3, 4, but uncommon (0–3%) in populations of European and African origins. Intermediate frequencies (30–50%) of the dry type are seen in populations of Southern Asia, the Pacific Islands, Central Asia and Asia Minor, as well as among the Native North American and Inuit of Asian ancestry2, 5. These figures show geographical gradient distributions. Earwax type may play a role in the axillary odor and possibly in breast cancer susceptibility6, although the association with breast cancer remains controversial7. Using a linkage analysis, we have previously assigned the earwax gene locus to the pericentromeric region of chromosome 16 (ref. 8).Distribution of allele A:

See also Gene for ear wax (BBC).
Nature Genetics (published online)
A SNP in the ABCC11 gene is the determinant of human earwax type
Koh-ichiro Yoshiura et al.
Human earwax consists of wet and dry types. Dry earwax is frequent in East Asians, whereas wet earwax is common in other populations. Here we show that a SNP, 538G right arrow A (rs17822931), in the ABCC11 gene is responsible for determination of earwax type. The AA genotype corresponds to dry earwax, and GA and GG to wet type. A 27-bp deletion in ABCC11 exon 29 was also found in a few individuals of Asian ancestry. A functional assay demonstrated that cells with allele A show a lower excretory activity for cGMP than those with allele G. The allele A frequency shows a north-south and east-west downward geographical gradient; worldwide, it is highest in Chinese and Koreans, and a common dry-type haplotype is retained among various ethnic populations. These suggest that the allele A arose in northeast Asia and thereafter spread through the world. The 538G right arrow A SNP is the first example of DNA polymorphism determining a visible genetic trait.
Link
DNA Testing: In Our Blood.

Newsweek has an article titled "DNA Testing: In Our Blood", which offers layman's introduction to the world of population genetics and geneaological testing.
From the article:
The research led Skorecki's team to Africa, where they tested members of the Lemba tribe, a group that believed they were descended from the Biblical land of Judea. Some of their DNA matched the Cohan signature. "We share a common paternal ancestry," says Skorecki. In 2001, Father Bill Sanchez, a Roman Catholic priest in Albuquerque, N.M., discovered he closely matched the Cohan signature, too. Sanchez's Jewish roots go back to Spain (his mother's heritage is Native American). Today he keeps pictures of his Christian and Jewish ancestors on his wall; in November he traveled to Israel. Now his niece Jessica Gonzales, 24, wants to go. Raised Catholic, she wants to learn more about her family roots. "I've been reading a lot about Judaism," she says.I don't know the specifics of Father Sanchez family origins, but I have to wonder when Skorecki et al. will finally decide to come clean about the significance of their "Cohen signature" and the little-known fact that it is neither a Jewish nor a Cohen-specific signature. I have a feeling that there are quite a lot of people out there who've been "reading about Judaism" based on false beliefs created by dubious science.
Also, from the article:
Last fall, Wells packed up 500 blood-collection tubes, needles, alcohol wipes and cheek swabs and headed off to Chad, one of the project's first testing sites, where he took 300 DNA samples from towns and villages around the country. Thirty-five to 40 came from members of the isolated Laal community, whose population, at fewer than 750, is declining. Wells fears that this community will die out within the next 10 to 30 years, taking with it valuable DNA and cultural traditions and an ancient language—information that could provide critical insights into the first people to live in Central Africa more than 40,000 years ago. "We can use DNA to figure out some of these great mysteries, to make sense of the past," says Wells.I guess I was right about Dr. Wells' activities in my New Year's predictions... and their name is Laal. Now, let's hope that Dr. Wells and his team will find some time to devote to the other 6,000,000,000-750 of us.
January 29, 2006
Y-chromosome analysis in the Near East

January 28, 2006
Dairying among European farmers
Dirty cooking pots dating to nearly 8,000 years ago reveal that some of Europe's earliest farming communities produced dairy products, such as cheese and yogurt.Antiquity 79(306): 882-894
Two separate studies indicate that Neolithic dairying took place in what are now Romania, Hungary and Switzerland.
The discoveries suggest people in these regions might have originally learned how to process milk-based foods from Asian farmers.
"From a diffusionist perspective, these findings lend support to the idea that the antiquity of dairying lies with the origins of animal domestication in southwest Asia some two millennia earlier, prior to its transmission to Europe in the seventh millennia B.C., rather than it being a later and entirely European innovation," wrote Oliver Craig, a scientist at Tor Vergata University in Rome, and colleagues in the first study published in the journal Antiquity.
Did the first farmers of central and eastern Europe produce dairy foods?
Oliver E. Craig et al.
Although the origins of domestic animals have been well-documented, it is unclear when livestock were first exploited for secondary products, such as milk. The analysis of remnant fats preserved in ceramic vessels from two agricultural sites in central and eastern Europe dating to the Early Neolithic (5900-5500 cal BC) are best explained by the presence of milk residues. On this basis, the authors suggest that dairying featured in early European farming economies. The evidence is evaluated in the light of analysis of faunal remains from this region to determine the scale of dairying. It is suggested that dairying — perhaps of sheep or goats — was initially practised on a small scale and was part of a broad mixed economy.
Link
Journal of Archaeological Science
Volume 33, Issue 1 , January 2006, Pages 1-13
Chemical analyses of organic residues in archaeological pottery from Arbon Bleiche 3, Switzerland – evidence for dairying in the late Neolithic
Jorge E. Spangenberg et al.
Fatty acids distribution and stable isotope ratios (bulk δ13C, δ15N and δ13C of individual fatty acids) of organic residues from 30 potsherds have been used to get further insights into the diet at the Late Neolithic (3384–3370 BC) site of Arbon Bleiche 3, Switzerland. The results are compared with modern equivalents of animal and vegetable fats, which may have been consumed in a mixed ecology community having agrarian, breeding, shepherd, gathering, hunting, and fishing activities. The used combined chemical and isotopic approach provides valuable information to complement archaeological indirect evidence about the dietary trends obtained from the analysis of faunal and plant remains. The small variations of the δ13C and δ15N values within the range expected for degraded animal and plant tissues, is consistent with the archaeological evidence of animals, whose subsistence was mainly based on C3 plants. The overall fatty acid composition and the stable carbon isotopic compositions of palmitic, stearic and oleic acids of the organic residues indicate that the studied Arbon Bleiche 3 sherds contain fat residues of plant and animal origin, most likely ruminant (bovine and ovine). In several vessels the presence of milk residues provides direct evidence for dairying during the late Neolithic in central Europe.
Link
January 27, 2006
Y-chromosome analysis in Turkey
UPGMA dendrogram based on all reported haplogroup frequencies:

PCA plot based on 10 most frequent haplogroups:

January 26, 2006
Greece and Italy Y-chromosome Analysis
UPGMA dendrogram:

PCA analysis:

Expanding Heads and Shrinking Faces
Researchers have found that the shape of the human skull has changed significantly over the past 650 years.The scientists took samples from a sunken ship, the Mary Rose, and from plague victims. The latter sample might be biased, since different head shapes have been shown in the past to have better chances of survival; however, the sample from the Mary Rose should be random, and the two historical samples tend to agree in their differentiation from modern populations.
Modern people possess less prominent features but higher foreheads than our medieval ancestors.
Writing in the British Dental Journal, the team took careful measurements of groups of skulls spanning across 30 generations.
The scientists said the differences between past and present skull shapes were "striking".
...
The two principal differences discovered were that our ancestors had more prominent features, but their cranial vault - the distance measured from the eyes to the top of the skull - was smaller.
Dr Peter Rock, lead author of the study and director of orthodontistry at Birmingham University, told the BBC News website: "The astonishing finding is the increased cranial vault heights.
"The increase is very considerable. For example, the vault height of the plague skulls were 80mm, and the modern ones were 95mm - that's in the order of 20% bigger, which is really rather a lot."
He suggests that the increase in size may be due to an increase in mental capacity over the ages.
From the paper:
For S-CVa, the measurement that is most representative of the anterior cranial fossa, there were significant differences between all three groups with size increasing through the ages. The anterior cranial fossa houses the frontal lobe of the brain, the great development of which is often held to be the major distinction between the human race and other primates. In particular the prefrontal areas are concerned with intellect35 and the increased intracranial dimensions and high foreheads of the modern group are evidence that brain size has increased over the centuries.British Dental Journal (2006); 200, 33-37. doi: 10.1038/sj.bdj.4813122Help
A cephalometric comparison of skulls from the fourteenth, sixteenth and twentieth centuries
W. P. Rock, A. M. Sabieha and R. I. W. Evans
Objectives To evaluate changes in the size and shape of the skull and jaws in British populations between the thirteenth and twentieth centuries.
Method Lateral cephalometric radiograms were obtained from skulls of three groups of subjects: 30 skulls were from the remains of those who died in the London Black Death epidemic of 1348, 54 skulls were recovered from the wreck of the Mary Rose which sank in 1545 and 31 skulls were representative of modern cephalometric values.
Results Horizontal measurements in the base of the anterior cranial fossa and in the maxillary complex were greater in the modern group than in the medieval skulls. Cranial vault measurements were significantly higher (P = 0.000) in the twentieth century skulls, especially in the anterior cranial fossa.
Conclusion Results suggest that our medieval ancestors had more prominent faces and smaller cranial vaults than modern man.
Link
January 25, 2006
Recent origins of East German population
In human populations, the correct historical interpretation of a genetic structure is often hampered by an almost inherent inability to differentiate between ancient and more recent influences upon extant gene pools.Let's hope that we'll see more papers brave and/or smart enough to look into recent causes of Y-chromosome distribution as opposed to yet-another-iteration of the Neolithic farmers vs. Paleolithic refugia explanations.
European Journal of Human Genetics (advance online publication)
Y-chromosomal STR haplotype analysis reveals surname-associated strata in the East-German population
Uta-Dorothee Immel et al.
Abstract
In human populations, the correct historical interpretation of a genetic structure is often hampered by an almost inherent inability to differentiate between ancient and more recent influences upon extant gene pools. One method to trace recent population movements is the analysis of surnames, which, at least in Central Europe, can be thought of as traits 'linked' to the Y chromosome. Illegitimacy, extramarital birth and changes of surnames may have substantially obscured this linkage. In order to assess the actual extent of correlation between surnames and Y-chromosomal haplotypes in Central Europe, we typed Y-chromosomal short tandem repeat markers in 419 German males from Halle. These individuals were subdivided into three groups according to the origin of their respective surname, namely German (G), Slavic (S) or 'Mixed' (M). The distribution of the haplotypes was compared by Analysis of Molecular Variance. While the M group was indistinguishable from group G (PhiST=-0.0008, P>0.5), a highly significant difference (PhiST=0.0277, P<0.001) was observed between the S group and the combined G+M group. This surprisingly strong differentiation is comparable to that of European populations of much larger geographic and linguistic difference. In view of the major migration from Slavic countries into Germany in the 19th century, it appears likely that the observed concurrence of Slavic surnames and Y chromosomes is of a recent rather than an early origin. Our results suggest that surnames may provide a simple means to stratify, and thereby to render more efficient, Y-chromosomal analyses of Central Europeans that target more ancient events.
Link
Origins of Faroe Islanders
Highly discrepant proportions of female and male Scandinavian and British Isles ancestry within the isolated population of the Faroe Islands
Thomas D Als et al.
Abstract
The Faroe Islands in the North Atlantic Ocean are inhabited by a small population, whose origin is thought to date back to the Viking Age. Historical, archaeological and linguistic evidence indicates that the present population of the Faroe Islands may have a mixture of Scandinavian and British Isles ancestry. In the present study we used 122 new and 19 previously published hypervariable region I sequences of the mitochondrial control region to analyse the genetic diversity of the Faroese population and compare it with other populations in the North Atlantic region. The analyses suggested that the Faroese mtDNA pool has been affected by genetic drift, and is among the most homogenous and isolated in the North Atlantic region. This will have implications for attempts to locate genes for complex disorders. To obtain estimates of Scandinavian vs British Isles ancestry proportions, we applied a frequency-based admixture approach taking private haplotypes into account by the use of phylogenetic information. While previous studies have suggested an excess of Scandinavian ancestry among the male settlers of the Faroe Islands, the current study indicates an excess of British Isles ancestry among the female settlers of the Faroe Islands. Compared to other admixed populations of the North Atlantic region, the population of the Faroe Islands appears to have the highest level of asymmetry in Scandinavian vs British Isles ancestry proportions among female and male settlers of the archipelago.
Link
How the steppe was colonized
Oxford Journal of Archaeology
Volume 24 Page 313 - November 2005
STEPS TO THE STEPPE: OR, HOW THE NORTH PONTIC REGION WAS COLONISED
IGOR MANZURA
Summary. The occupation of the steppe region north of the Black Sea by farming or herding groups in the fifth and fourth millennia BC has been a controversial question. At the core of the problem is the changing relationship between Cucuteni-Tripole farming groups in the forest-steppe zone and their neighbours in the true steppe zone. Three phases of this relationship are discussed, in the Early Copper Age, Late Copper Age and Early Bronze Age (c.5000–3000 BC), during which different forms of exchange and acculturation took place, each with its own social and economic characteristics. The role of environmental change, and the significance of burial monuments in the process of cultural convergence, are evaluated. The process is discussed both in terms of general models of social transformation, and by comparison with other areas of Europe where similar processes of interaction were taking place.
Link
January 24, 2006
Polynesian inflow into Bismarck archipelago Melanesians
(Early View)
Brief communication: Mitochondrial DNA variation suggests extensive gene flow from Polynesian ancestors to indigenous Melanesians in the northwestern Bismarck Archipelago
Jun Ohashi et al.
Abstract
Archaeological, linguistic, and genetic studies show that Austronesian (AN)-speaking Polynesian ancestors came from Asia/Taiwan to the Bismarck Archipelago in Near Oceania more than 3,600 years ago, and then expanded into Remote Oceania. However, it remains unclear whether they extensively mixed with indigenous Melanesians who had populated the Bismarck Archipelago before their arrival. To examine the extent of admixture between Polynesian ancestors and indigenous Melanesians, mitochondrial DNA (mtDNA) variations in the D-loop region and the cytochrome oxidase and lysine transfer RNA (COII/tRNALys) intergenic 9-bp deletion were analyzed in the following three Oceanian populations: 1) Balopa Islanders as AN-speaking Melanesians living in the northwestern end of the Bismarck Archipelago, 2) Tongans as AN-speaking Polynesians, and 3) Gidra as non-Austronesian-speaking Melanesians in the southwestern lowlands of Papua New Guinea. Phylogenetic analysis of mtDNA sequences revealed that more than 60% of mtDNA sequences in the Balopa Islanders were very similar to those in Tongans, suggesting an extensive gene flow from Polynesian ancestors to indigenous Melanesians. Furthermore, analysis of pairwise difference distributions for the D-loop sequences with the 9-bp deletion and the Polynesian motif (i.e., T16217C, A16247G, and C16261T) suggested that the expansion of Polynesian ancestors possessing these variations occurred approximately 7,000 years ago.
Link
Balkan Y-chromosome analysis
January 23, 2006
Criticism on papers regarding Jewish genetics
UPDATE
The point of the article is that the frequency of the LMH in the original publication is much less than that in the later publication. In the initial publication it is 21% among Ashkenazi Levites and it becomes 42% in the later publication.
So, either:
1. Significant typing errors occurred in the first publication, and the 21% frequency is widely off-the-mark. Did they publish an erratum, a letter to the editor, an announcement on their website, correcting this error? Or did they allow their result to stand in the scientific literature, even though they knew it had serious errors?
or:
2. They used a biased convenience sample in their second article which just so happened to have twice the frequency of the LMH?
So, either the researchers botched the first publication and kept quiet about their mistake, or they massaged their second sample to increase the frequency of the LMH.
I would be interested in any alternative interpretations.
HOMO - Journal of Comparative Human Biology
(Article in Press)
Ashkenazi levites’ “Y modal haplotype” (LMH) – an artificially created phenomenon?
A. Zoossmann-Diskin
Abstract
The article on the Y chromosomes of Ashkenazi Levites (Behar et al., 2003. Am. J. Hum. Genet. 73, 768–779) is the fourth in a series on the Y chromosomes of the three Jewish male castes: Cohanim (priests), Levites (priests’ helpers) and Israelites (lay people). It became apparent that there is a problem with omission of samples when the second article “Origins of Old Testament priests” (Thomas et al., 1998. Nature 394, 138–140) was published. In the fourth article a remarkable 55% of the Ashkenazi Levite samples from the earlier 1998 study are not included. This causes the “Levite modal haplotype” to double its frequency from 21% of the Ashkenazi Levite sample in 1998 to 42% of the Ashkenazi Levite sample in 2003. The authors offer three main explanations:
(1) The studies are independent using different sample sets.
(2) Typing errors and poor quality exclude samples from future studies.
(3) Correction of typing errors means that some samples are classified under different haplotypes.
The explanations offered to the problem of omitting samples from subsequent studies after their haplotypes or partial haplotypes are known, are not convincing. Consequently their sample sets cannot be considered random and non-biased. At the least, these laboratories have bad practices of sample handling and many typing errors, which are enough to invalidate their studies.
Link
January 22, 2006
Ancient Basques were not isolated (mtDNA)
And an exciting hint at future developments in ancient DNA:
Furthermore, two individuals in a group burial in the SE revealed the existence of a mutation in the Y chromosome (haplogroup R1b3b; unpublished data), considered to originate in the Basque Country, where it now presents the highest frequency in Europe (7.1%, as opposed to 0.9% in the Iberian Peninsula; Alonso et al., 2005).
Am J Phys Anthropol. 2006 Jan 19; [Epub ahead of print]
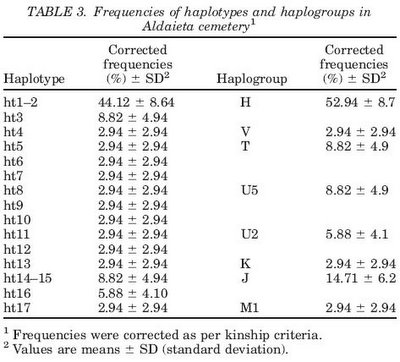
Insights into the "isolation" of the Basques: mtDNA lineages from the historical site of Aldaieta (6th-7th centuries AD).
Alzualde A, Izagirre N, Alonso S, Alonso A, Albarran C, Azkarate A, de la Rua C.
We analyzed the hypervariable region I (HVR-I) sequence variability of the mitochondrial DNA (mtDNA) of individuals buried at Aldaieta (6th-7th centuries AD) in order to find out more about the biosocial implications of this cemetery. The results, fully authenticated by means of diverse criteria (analysis of duplicates, replication in an independent laboratory, quantification of target DNA, and sequencing and cloning of polymerase chain reaction products), suggest that Aldaieta largely consists of autochthonous individuals who shared common funereal customs with the late Ancient North Pyrenean cemeteries of Western Europe (the Reihengraberfelder), a cultural influence possibly accompanied by a certain genetic flow. Furthermore, the distribution of mtDNA lineages in the cemetery highlighted the existence of a significant number of family relationships, supporting the belief that it was a stable settlement and not a group that had haphazardly settled in the area. Finally, this paper stresses the importance of ancient DNA data for reconstructing the biological history of human populations, rendering it possible to verify certain hypotheses based solely on current population data. The presence at Aldaieta of an mtDNA lineage originating in Northwest Africa testifies to the existence of contact between the Iberian Peninsula and Northwest Africa prior to the Moorish occupation. Both this latter discovery and the high frequency of haplogroup J at the Aldaieta cemetery raise questions about the generally accepted belief that, since ancient times, the influence of other human groups has been very scarce in the Basque Country.
Link
Tyrolean Iceman had mtDNA haplogroup K1
Fine characterization of the Iceman's mtDNA haplogroup.
Rollo F, Ermini L, Luciani S, Marota I, Olivieri C, Luiselli D.
Starting from specimens of the intestinal contents of the so-called Tyrolean Iceman or Otzi (5,350-5,100 years before present), it was possible by polymerase chain reaction to amplify fragments of the human mitochondrial DNA (mtDNA) control region that correspond to the sequence found in 1994 at the Munich and Oxford laboratories and which had been attributed to the original DNA of the mummy. The particularly favorable condition of the specimens, showing very low contamination levels, made it easier to extend the analyses to the coding region, which had not previously been considered. The mtDNA of the European population is currently divided into nine (H, T, U, V, W, X, I, J, and K) main groups (haplogroups). The K haplogroup, in particular, is composed of two (K1 and K2) subclusters. The results demonstrate that the Iceman's mtDNA belongs to the K1 subcluster, yet it does not fit any of the three known branches (a, b, and c) into which the K1 subcluster is presently divided. In addition, some other sites, reported to be linked to environmental adaptation or pathologies, were investigated.
Link
DNA examination of ancient dental pulp incriminates typhoid fever as a probable cause of the Plague of Athens.
DNA examination of ancient dental pulp incriminates typhoid fever as a probable cause of the Plague of Athens.
Papagrigorakis MJ, Yapijakis C, Synodinos PN, Baziotopoulou-Valavani E.
BACKGROUND: Until now, in the absence of direct microbiological evidence, the cause of the Plague of Athens has remained a matter of debate among scientists who have relied exclusively on Thucydides' narrations to introduce several possible diagnoses. A mass burial pit, unearthed in the Kerameikos ancient cemetery of Athens and dated back to the time of the plague outbreak (around 430 BC), has provided the required skeletal material for the investigation of ancient microbial DNA. OBJECTIVE: To determine the probable cause of the Plague of Athens. METHOD: Dental pulp was our material of choice, since it has been proved to be an ideal DNA source of ancient septicemic microorganisms through its good vascularization, durability and natural sterility. RESULTS: Six DNA amplifications targeted at genomic parts of the agents of plague (Yersinia pestis), typhus (Rickettsia prowazekii), anthrax (Bacillus anthracis), tuberculosis (Mycobacterium tuberculosis), cowpox (cowpox virus) and cat-scratch disease (Bartonella henselae) failed to yield any product in 'suicide' reactions of DNA samples isolated from three ancient teeth. On the seventh such attempt, DNA sequences of Salmonella enterica serovar Typhi were identified providing clear evidence for the presence of that microorganism in the dental pulp of teeth recovered from the Kerameikos mass grave. CONCLUSION: The results of this study clearly implicate typhoid fever as a probable cause of the Plague of Athens.
Link
Secret of Athens plague unravelled
Secret of ancient Athens plague is being unraveled
Greek scientists find typhoid after excavating graves
Kerameikos, Athens’s ancient cemetery, has yielded conclusive evidence as to the nature of the plague that decimated a third of the population of the ancient city and influenced the outcome of the Peloponnesian Wars. Scientists at Athens University’s School of Dentistry have used molecular biology to help solve the riddle of one of history’s biggest mysteries.
By Dr Manolis Papagrigorakis (1)
Recent findings from a mass grave in the Ancient Cemetery of Kerameikos in central Athens show typhoid fever may have caused the plague of Athens, ending centuries of speculation about what kind of disease killed a third of the city’s population and contributed to the end of its Golden Age.
Examined by a group of Greek scientists coordinated by Dr Manolis Papagrigorakis of Athens University’s School of Dentistry, the findings provide clear evidence that Salmonella enterica serovar Typhi was present in the dental pulp of teeth recovered in remains from the mass grave.
The plague that decimated the population of Athens in 430-426 BC was a deciding factor in the outcome of the Peloponnesian Wars, ending the Golden Age of Pericles and Athens’s predominance in the Mediterranean.
It broke out during the siege of the city by the Spartans in the early summer of 430 BC; after a brief hiatus in 428 BC, the epidemic returned in the winter of 427 BC and lasted until the winter of the following year. It is assumed that one-third of the Athenians, including one-fourth of their army and their charismatic leader, Pericles, perished in the epidemic.
All data pertaining to the disease’s outbreak and its clinical characteristics were until now based on the account by the fifth-century-BC Greek historian Thucydides, who himself fell ill with the plague but recovered. In his famous history of the Peloponnesian Wars, Thucydides gives detailed descriptions that have formed the basis of several hypotheses regarding its nature. However, researchers had never managed to agree on the identity of the plague due to the lack of definite microbiological proof in the absence of paleopathologic evidence. Several pathogens have been putatively implicated in the emergence and spreading of the disease.
In recent decades, molecular biology tools (DNA PCR and sequencing techniques) have made it possible to detect and, furthermore, specifically identify microbial DNA fragments in ancient human skeletal remains, thus making possible the retrospective diagnoses of ancient diseases.
In 1994-1995, under the supervision of archaeologist Effi Baziotopoulou-Valavani for the Fourth Prehistoric and Classical Antiquities Ephorate, excavations of a mass burial pit unearthed in the Ancient Cemetery of Kerameikos in Athens provided the required skeletal material for the investigation of ancient microbial DNA.
The grave yielded the remains of about 150 individuals and were dated, through archaeological site documentation, to around the time of the plague outburst between 430-426 BC. The remains were found piled up in a manner that indicated a hasty burial without the usual care dictated by the respect that ancient Greeks usually showed for the dead.
Dental pulp was the material of choice in this research, as its good vascularization, durability and natural sterility has proven to be an ideal source of ancient DNA, also providing for the recovery of adequate genetic material of specific septicemic microorganisms which after death remain trapped in the dental pulp and become mummified.
Using modern laboratory methods under strict sterile conditions at the molecular neurobiology laboratory at Athens University’s medical school, the research team first found the existence of microbial DNA in the dental pulp. This DNA was then separated and subjected to successive tests to identify which of the possible microbes was linked in the past with the Athens plague.
Teeth from three different skeletons were examined. After six negative results from six candidate microbes, a positive reaction was found for Salomonella enterica serovar Typhi, which is responsible for the appearance of typhoid fever.
The correspondence with the genes examined in the ancient DNA with known sequences of the contemporary form of the microbe was as high as 99 percent.
This evidence allowed for a definite conclusion regarding the microbes found in the teeth of the three bodies from the mass burial pit — the presumed victims of the Athens plague.
Typhoid fever almost certainly played a part in causing the Athens plague, either exclusively or in combination with another — and so far unknown — infection.
Even today, typhoid fever is a major health problem on a global scale. Every year there are about 20 million new cases that lead to about 600,000 deaths in the developing world where overpopulation, inadequate water supplies and hygiene, as well as poor access to health services, allow epidemics to spread with tragic results.
Overcrowding and resultant public health problems — as well as standards of personal hygiene — in the besieged city of Athens in 430 BC as described by Thucydides would have been sufficient to allow the disease to appear and then develop into a deadly epidemic.
The scientifically documented diagnosis of typhoid fever is in accordance with many of the clinical characteristics of the Athens plague as described by Thucydides. The differences in the modern form of the disease from Thucydides’ references pose another challenge for the Greek research team.
Studying the historical aspects of infectious diseases can be a powerful tool for several disciplines to learn from. We believe this report to be of outstanding importance for many scientific fields, since it sheds light on one of the most debated enigmas in medical history. Archaeology, paleontology, history, paleopathology, certain fields of medicine, anthropology and even genetics, molecular biology and studies on evolution are clearly implicated in such matters and can benefit from relevant studies.
The results of this particular study are extremely important as they shed light on one of the greatest mysteries in world history. Also important is the fact that the research was organized, carried out and completed by Greek scientists at Greek research centers, under the aegis of Athens University.
(1) Dr Papagrigorakis is an assistant professor at Athens University’s School of Dentistry.
The other authors of the study, published today in the International Journal of Infectious Diseases, are geneticist Christos Yiapitzakis, orthodontist Philippos Synodinos and archaeologist Effi Baziotopoulou-Valavani.
January 20, 2006
Sexual dimorphism in gene expression in human brain
Most notably, in man the brain is sexually dimorphic with respect to asymmetry (the cerebral torque) from right frontal to left occipital across the anteroposterior axis (Barrick et al. 2004). Females are more strongly right-handed than males, acquire words more rapidly (Crow et al. 1998) and have faster brain growth (Kretschmann et al. 1979). By contrast, adult brain size is greater in males than females, cerebral asymmetry is more marked, particularly in the posterior part of the brain (Barrick et al. 2004) and there is a modest male superiority for spatial ability. Although most sexual dimorphisms are caused by male and female gonadal secretions, there is evidence for a direct role of sex chromosomes genes in brain sexual differentiation (Carruth et al. 2002; reviewed in Arnold 2004). In this context, an imbalance of PCDH11X/Y products between sexes might be relevant considering that these genes are expressed from early in development (Blanco et al. 2000).It's nice to see that some women scientists demonstrate their excellence by writing excellent papers about male-female biological differences rather than engage in knee-jerk reactions to legitimate scientific speculation.
Molecular Biology and Evolution (online first)
Inactivation status of PCDH11X: sexual dimorphisms in gene expression levels in brain
Alexandra M. Lopes et al.
Abstract Genes escaping X-inactivation are predicted to contribute to differences in gene dosage between the sexes and are the prime candidates for being involved in the phenotype observed in individuals with X chromosome aneuploidies. Of particular interest is ProtocadherinX (PCDH11X or PCDHX), a recently described gene expressed in brain. In humans, PCDH11X has a homologue on the Y chromosome and is predicted to escape from X-inactivation. Employing bisulphite sequencing analysis we found absence of CpG island methylation on both the active and the inactive X chromosomes, providing a strong indication that PCDH11X escapes inactivation in humans. Furthermore, a sexual dimorphism in levels of expression in brain tissue was observed by quantitative real-time PCR, with females presenting an up to 2-fold excess in the abundance of PCDH11X transcripts. We relate these findings to sexually dimorphic traits in the human brain. Interestingly, PCDH11X/Y gene pair is unique to Homo sapiens, since the X-linked gene was transposed to the Y chromosome after the human–chimpanzee lineages split. Although no differences in promoter methylation were found between humans and chimpanzees, evidence of an upregulation of PCDH11X in humans deserves further investigation.
Link
January 19, 2006
Amazonian Hunter-Gatherers grasp geometry
Science 20 January 2006: 381-384
Core Knowledge of Geometry in an Amazonian Indigene Group
Stanislas Dehaene, Véronique Izard, Pierre Pica, and Elizabeth Spelke
Does geometry constitute a core set of intuitions present in all humans, regardless of their language or schooling? We used two nonverbal tests to probe the conceptual primitives of geometry in the Mundurukú, an isolated Amazonian indigene group. Mundurukú children and adults spontaneously made use of basic geometric concepts such as points, lines, parallelism, or right angles to detect intruders in simple pictures, and they used distance, angle, and sense relationships in geometrical maps to locate hidden objects. Our results provide evidence for geometrical intuitions in the absence of schooling, experience with graphic symbols or maps, or a rich language of geometrical terms.
Link
Mesolithic of Baltic Sea basin
Mobility, contact, and exchange in the Baltic Sea basin 6000–2000 BC
Marek Zvelebil
My intention in this paper is to outline the main features and principal aspects of contact and exchange among the later prehistoric hunter–gatherers (late Mesolithic and post-Mesolithic) in the Baltic Sea basin, which covers the southern and eastern reaches of Northern Europe, and to summarise the main advances in current research. The area broadly covered includes the Baltic Sea basin that has provided effective routes for communication between the coastal regions surrounding the Baltic Sea, central Baltic islands, and regions further away in the north European Plain, inland regions of Fennoscandia and Russia that could be reached by an extensive network of major rivers and lakes. Effective transport for negotiating these routes both in the summer and winter existed already from the early Mesolithic. Goods moved along these routes included a wide range of artefacts discussed in the paper. Geographically, exchange was organised at three levels: regionally, inter-regionally, and over long distances. Each mode of exchange was probably organised along different lines socially, and each served to implement wide-ranging social strategies for the general purposes of social reproduction, mate exchange and biological reproduction, as well as the spread of innovations. In the concluding section, I discuss the nature of contacts and consequences of exchanges between the early farming communities and the hunter–gathering groups within the framework of the core-periphery relations.
Link
The face of Agamemnon

Dickinson's ingenuous argument doesn't address the claims about the authenticity of the above mask -although he demonstrates how difficult it would be for Schliemann to carry out the alleged fraud. Rather, he notes a telegram sent by S. whose contents are incompatible with the idea that NM624 was considered by him to be Agamemnon.
"“In the last tomb three bodies, one without ornaments. Have telegraphed to
Nauplia for a painter, to preserve the dead man with the round face [my
italics]. This one is very like the picture which my imagination formed of
Agamemnon long ago.”7 Surely, the “round-faced man” could refer only to NM 623."

NM 623, pictured above was also extensively described by S. in his book, and was associated with richer burial goods than NM 624, thus making it more likely that he would be the high king associated by S. with Agamemnon.
Of course, we now know that the Shaft Graves date several centuries before the Trojan War, so none of the buried individuals is really Agamemnon.
Link (pdf) Hesperia Volume 74, Number 3, October 2005.
The fate of the well-groomed Irish bog men
A team lead by researchers at the National Museum of Ireland studied the two bodies. The scientists say the fingerprint whorls of Oldcroghan man are as clear as any living person's.
"He had very well manicured nails, and his fingertips and hands were indicative of somebody who didn't carry out any manual labor. So we presume he came from the upper echelons of society," said Isabella Mulhall, the museum's Bog Bodies Project coordinator.
...
Had the two bog men met, Oldcroghan man [DP: 198cm] would have towered over Clonycavan man, who measured just 5 feet, 2 inches (157 centimeters) tall.
Perhaps to compensate for his short stature, Clonycavan man coiffed up his hair using an early hair gel.
"Naturally enough, he wanted to make himself look grander," Mulhall said. "It's a bit like someone wearing platform shoes."
...
While both bog men appeared to be aristocratic dandies of their day, they still met horrible deaths.
Oldcroghan man shows signs of cruel torture before he was beheaded.
"He was stabbed, his nipples were sliced, and he had holes cut in his upper arms through which a rope was threaded in order to restrain him," Mulhall said. He was also cut in half across the torso.
Meanwhile, Clonycavan man suffered three axe blows to the head, plus one to his chest and was also disemboweled.
"There was definitely an attempt to use several different methods to traumatize and torture the men," Mulhall added.
January 18, 2006
Flawed mtDNA data sets
It is important to critically evaluate published data sets, because otherwise data sets, even as flawed as the one from Nasidze & Stoneking (2001), may become innocently incorporated in further data analyses by other authors, yielding therefore a growing avalanche of a priori flawed results (e.g., Bulayeva et al. 2003; Vernesi et al. 2004). The lessons to be learnt from this case are the following:
1. Researchers should not rush into publication with poor quality DNA sequences and without checking for artificial patterns in the data.
2. Any major inference from mtDNA data must be accompanied by the primary data, which should be submitted to GenBank before submission (and made available there as soon as the paper is published electronically), and additionally displayed in the paper or as an electronic supplement (either in the form of a diagram or a table) in order to enable quick evaluation by referees and readers.
3. Major analyses carried out and interpreted in the paper should be displayed in the form of tables and diagrams.
4. Referees of a submitted manuscript should routinely request additional data and details about analyses that are necessary to check results fully (although this may be difficult to achieve in view of the overabundance of mtDNA papers submitted to journals).
It is nice to see that some researchers take the time to critically evaluate existing results and methodologies.
Ann Hum Genet (early view)
Quality Assessment of DNA Sequence Data: Autopsy of A Mis-Sequenced mtDNA Population Sample
H.-J. Bandelt, and T. Kivisild
Published DNA data sets constitute a body of sequencing results resting in silico that are supposed to reflect the variation of (once) living cells. In cases where the DNA variation reported is suspected to be fraught with artefacts, an autopsy of the full body of data is needed to clarify the amount and causes of mis-sequencing. In this paper we elaborate on strategies that allow a clear-cut identification of the problems in severely flawed mtDNA data. This approach is applied, by way of example, to a data set of HVS-I sequences from the Caucasus, published by Nasidze & Stoneking in 2001. These data bear numerous ambiguous nucleotide positions and suffer from an even higher number of phantom mutations, indicating that severe biochemical problems adversely influenced those sequencing results at the time. Furthermore, systematic omission of sequences with a long C-stretch (incurred by a transition at position 16189) must have severely biased the data set. Since no complete correction of these data has appeared to date, this example of mis-sequencing necessitates circumstantial evidence that is bullet-proof.
Link
Self-defined race/ethnicity and true underlying genetic structure
For the three racial/ethnic groups combined, the posterior probabilities of K obtained from Structure, Pr(X | K), were close to 1 when K = 3 and close to 0 for all other tested K values. This means that the optimal number of clusters was three.69% of skin color variation could be explained by self-reported ancestry and there were 10 mis-assigned individuals out of 1,334. 4 of them were likely of mixed recent ethnicity. Additionally, African Americans, and Hispanics had a higher variance in skin color, as can be expected due to their multi-racial origins.
Ann Hum Genet (early view)
The role of Self-Defined Race/Ethnicity in Population Structure Control
X-Q. Liu et al.
Population-based association studies are powerful tools for the genetic mapping of complex diseases. However, this method is sensitive to potential confounding by population structure. While statistical methods that use genetic markers to detect and control for population structure have been the focus of current literature, the utility of self-defined race/ethnicity in controlling for population structure has been controversial. In this study of 1334 individuals, who self-identified as either African American, European American or Hispanic, we demonstrated that when the true underlying genetic structure and the self-defined racial/ethnic groups were roughly in agreement with each other, the self-defined race/ethnicity information was useful in the control of population structure.
Link
Chimpanzees aren't altruistic
In Jensen's study, chimpanzees from the Wolfgang Koehler Primate Research Centre in Leipzig were given a choice; by pulling on a rope they could either deliver food to another chimpanzee or they could deliver it to an empty room. In both cases, the chimpanzee pulling the rope did not receive any food itself. Contrary to initial expectations the chimpanzees behaved neither altruistic nor spiteful. According to the researchers, both characteristics therefore seem to be human-specific.
An altruistic chimpanzee would give food to its neighbour, despite the effort in pulling the food, and a spiteful chimpanzee would prevent its neighbour from having the food by delivering it to the empty room.
'I predicted chimps would be spiteful. I thought if they knew they couldn't have the food, they wouldn't let anyone else have it.' Jensen found that half the time, the chimpanzees did nothing. A quarter of the time they delivered food to their neighbour, then a quarter of the time to the empty room. This demonstrated neither altruism nor spite.
'They didn't seem to care about the other guy one way or the other. All that concerned them was getting the food and they were completely focused on that. Even when they knew they couldn't have the food, they didn't help the other chimp but they weren't spiteful either.'
In contrast, humans are obviously altruistic. We give blood, we donate money to charity, and we volunteer to help strangers. This kind of altruism has never been demonstrated in any other animal except for humans and some believe it is one of the characteristics that makes us human. But Jensen says spite is just as important. As a form of punishment, spite can encourage cooperative behaviour by penalising cheaters.
'Punishing others is usually costly to yourself, whether that's the taxpayer or the lawmakers but punishment is still a natural part of modern society. We punish theft, murder and countless other crimes to keep the fabric of society together. Perhaps human society is where it is today because spite exists and there is a mechanism to punish cheaters.'
If altruism and spite are unique to humans and are not present in chimpanzees, then it is likely that these characteristics have arisen in the last 6 million years since humans and chimpanzees shared a common ancestor. Humans' intense regard for each other, either positive or negative, may have made an important contribution to our ability to cooperate, our sense of fairness, and the morality that defines today's society.
January 17, 2006
Some more random (?) facts
"The Lycians are in good truth anciently from Crete; which island, in former days, was wholly peopled with barbarians. A quarrel arising there between the two sons of Europa, Sarpedon and Minos, as to which of them should be king, Minos, whose party prevailed, drove Sarpedon and his followers into banishment. The exiles sailed to Asia, and landed on the Milyan territory. Milyas was the ancient name of the country now inhabited by the Lycians: the Milyae of the present day were, in those times, called Solymi. So long as Sarpedon reigned, his followers kept the name which they brought with them from Crete, and were called Termilae, as the Lycians still are by those who live in their neighbourhood. But after Lycus, the son of Pandion, banished from Athens by his brother Aegeus had found a refuge with Sarpedon in the country of these Termilae, they came, in course of time, to be called from him Lycians. Their customs are partly Cretan, partly Carian. They have, however, one singular custom in which they differ from every other nation in the world. They take the mother's and not the father's name. Ask a Lycian who he is, and he answers by giving his own name, that of his mother, and so on in the female line. Moreover, if a free woman marry a man who is a slave, their children are full citizens; but if a free man marry a foreign woman, or live with a concubine, even though he be the first person in the State, the children forfeit all the rights of citizenship."
Herodotus, Histories, 1.173
"Lycian, a language of SW Anatolia for which there are written records dated from about the 5th to 4th cent. B.C., may have been a continuation of Luwian."
Anatolian Languages in The Columbia Encyclopedia
"As in the Bronze Age syllabary more certainly evolved for the recording of Luvian, the so-called Hittite hieroglyphs, the sound values of the Linear A signary seem designed to represent exactly the Luvian phonetic system having 3 long vowels, 3 short. And like its Anatolian counterpart, where certain of the syllabograms demonstrably match the sound of the first syllable in Luvian of the object that inspired its sign-shapes, the Linear A signary too would appear to have been built on the acrophonic principle. So far, some three dozen matches between Ventris' Linear B sound values (slightly altered to reflect the Greek 5-vowel system) and the Luvian name for the object often still identifiable in the Linear A sign--e.g., AB40/L28: WI as in WIdula-, the Hittite (and likely Luvian) term for 'chariot box'."
Edwin L. Brown faculty page
Neandertals may have arrived in Europe by sea
Spanish investigators believe they may have found proof that neanderthal man reached Europe from Africa not just via the Middle East but by sailing, swimming or floating across the Strait of Gibraltar.
Prehistoric remains of hunter-gatherer communities found at a site known as La Cabililla de Benzú, in the Spanish north African enclave of Ceuta, are remarkably similar to those found in southern Spain, investigators said. Stone tools at the site correspond to the middle palaeolithic period, when neanderthal man emerged, and resemble those found across Spain.
"This could break the paradigm of most investigators, who have refused to believe in any contact in the palaeolithic era between southern Europe and northern Africa," investigator José Ramos explained in the University of Cadiz's research journal.
Although the scientists have not yet reached definite conclusions, they say the evidence that neanderthal man mastered some primitive techniques for crossing the sea into Europe from the coast near Ceuta looks promising.
...
Fauna and flora evidence from the same era suggested both sides of the Mediterranean were by no means isolated. A neanderthal ability to travel across small stretches of sea would help explain why the Iberian peninsula has older examples of human remains than, say, France.
Mr Ramos said: "If the only way of getting to Europe was via the Middle East then, theoretically, they should have got to France before reaching Spain."
January 16, 2006
Columbus and Colom
DNA may expose the shady origins of Columbus (excerpt)
About 120 Catalans are to donate samples of their saliva this week to a team of geneticists headed by José Antonio Lorente Acosta, the head of the laboratory of genetic identification at Granada University.
Tests on another 180 people sharing the name Colom will follow in Mallorca and Valencia. Investigators will compare the results with the DNA from Columbus's illegitimate son, Hernando, whose remains lie in Seville Cathedral.
"We're not looking for descendants of Columbus but a common ancestor who may be the link between the admiral and today's Coloms. If we find a Y chromosome (the only one that males inherit by the paternal line), we could say they were related," a spokesperson for Acosta said this week.
January 13, 2006
Sahoo et al. (2006) online (Indian Y chromosome variation)
Interestingly Sanghamitra Sahoo seems to have published a paper on the same topic only two months after Sanghamitra Sengupta did.
UPDATE
It is unfortunate that this paper uses a limited number of UEP markers. Hopefully, future studies will start to seek and test more recently derived markers, which are the only ones that can really address recent events authoritatively.
Moreover, no STR markers were typed, thus further limiting any possible inferences about the time depth of the various Indian lineages.
A real problem with the study is that it performed an "admixture analysis" which considered the modern Central Asians as representative of the prehistoric ones. As it is well known, Central Asians of today have substantial Mongoloid admixture from the proto-historical and historical period and are not representative of the ancient Indo-Iranian groups of the steppe.
In any case, the observations of the authors about the distribution of haplogroups in India are broadly similar to those of the other recent study, and we have to agree that the wholesale assignment of J/R/L Y chromosomes to a recent invasion cannot really be sustained.
From the paper's conclusions:
It is not necessary, based on the current evidence, to look beyond South Asia for the origins of the paternal heritage of the majority of Indians at the time of the onset of settled agriculture. The perennial concept of people, language, and agriculture arriving to India together through the northwest corridor does not hold up to close scrutiny. Recent claims for a linkage of haplogroups J2, L, R1a, and R2 with a contemporaneous origin for the majority of the Indian castes’ paternal lineages from outside the subcontinent are rejected, although our findings do support a local origin of haplogroups F* and H. Of the others, only J2 indicates an unambiguous recent external contribution, from West Asia rather than Central Asia. The current distributions of haplogroup frequencies are, with the exception of the O lineages, predominantly driven by geographical, rather than cultural determinants. Ironically, it is in the northeast of India, among the TB groups that there is clear-cut evidence for large-scale demic diffusion traceable by genes, culture, and language, but apparently not by agriculture.This certainly seems reasonable. J2 is largely restricted to the upper castes in India, and its young age is very suggestive of an external arrival in Neolithic and later times. It is certainly beginning to stand out as the most important exogenous genetic component in the Indian population.
The conclusion that J2 arrived in India from West and not Central Asia is not well-founded, because in the Middle East J*(xJ2) is frequent, but completely lacking in India. But, perhaps, most of Middle Eastern J*(xJ2) expanded recently, with the growth of the Semitic groups and was not present in the parental population from which Indian J2 is derived. Central Asia cannot be rejected so easily though as a source for Indian J2, because the present-day Central Asians have components (within N/C/O/Q) which were probably added by Mongoloid groups recently, and are not representative of the prehistoric populations.
The next step should be to develop informative recent markers in haplogroups J2a and R1a1 to finally establish whether some of these can be unambiguously related to particular Western Eurasian populations. This might be the decisive step to conclude whether Renfrew's hypothesis A (arrival of IE languages to India with early farmers) or hypothesis B (arrival of IE languages to India with IE-ized pastoral nomads) is the correct one.
PNAS (online early)
A prehistory of Indian Y chromosomes: Evaluating demic diffusion scenarios
Sanghamitra Sahoo et al.
Understanding the genetic origins and demographic history of Indian populations is important both for questions concerning the early settlement of Eurasia and more recent events, including the appearance of Indo-Aryan languages and settled agriculture in the subcontinent. Although there is general agreement that Indian caste and tribal populations share a common late Pleistocene maternal ancestry in India, some studies of the Y-chromosome markers have suggested a recent, substantial incursion from Central or West Eurasia. To investigate the origin of paternal lineages of Indian populations, 936 Y chromosomes, representing 32 tribal and 45 caste groups from all four major linguistic groups of India, were analyzed for 38 single-nucleotide polymorphic markers. Phylogeography of the major Y-chromosomal haplogroups in India, genetic distance, and admixture analyses all indicate that the recent external contribution to Dravidian- and Hindi-speaking caste groups has been low. The sharing of some Y-chromosomal haplogroups between Indian and Central Asian populations is most parsimoniously explained by a deep, common ancestry between the two regions, with diffusion of some Indian-specific lineages northward. The Y-chromosomal data consistently suggest a largely South Asian origin for Indian caste communities and therefore argue against any major influx, from regions north and west of India, of people associated either with the development of agriculture or the spread of the Indo-Aryan language family. The dyadic Y-chromosome composition of Tibeto-Burman speakers of India, however, can be attributed to a recent demographic process, which appears to have absorbed and overlain populations who previously spoke Austro-Asiatic languages.
Link
Prostate and renal cancer and mtDNA haplogroup U
This finding stresses that lineage-specific selection may be operating in the human mitochondrial genome. However, this is unlikely to affect the entire haplogroup U, which is the oldest lineage found in Caucasoids and occurs at non-trivial frequencies.
J Urol. 2006 Feb;175(2):468-73.
North american white mitochondrial haplogroups in prostate and renal cancer.
Booker LM, Habermacher GM, Jessie BC, Sun QC, Baumann AK, Amin M, Lim SD, Fernandez-Golarz C, Lyles RH, Brown MD, Marshall FF, Petros JA.
PURPOSE: While the mitochondrion is known to be a key mediator of apoptosis, there has been little inquiry into the inheritance pattern of mitochondria in patients with cancer. We compared the mtDNA haplotype in patients with prostate and renal cancer to that in controls to determine if there is an association between mitochondrial genotype and cancer. MATERIALS AND METHODS: Haplotyping was performed using polymerase chain reaction/digest identification of key polymorphic sites in the mitochondrial genome. A total of 121 and 221 white men with renal and prostate cancer, respectively, were identified following pathological confirmation of cancer, while 246 white controls were selected randomly from a bank of cadaveric organ donor DNA. Statistical analysis was performed and ORs were calculated. RESULTS: Mitochondrial haplogroup U was a highly significant risk factor for prostate and renal cancer vs controls (16.74% and 20.66% vs 9.35%, Fisher's exact test p = 0.019 and 0.005, respectively). The association remained statistically significant in renal cancer even after Bonferroni adjustment for multiple comparisons. Haplogroup U carried an OR of 1.95 for prostate cancer and an OR of 2.52 for renal cancer. CONCLUSIONS: The inheritance of mitochondrial haplogroup U is associated with an approximately 2-fold increased risk of prostate cancer and 2.5-fold increased risk of renal cancer in white North American individuals. Therefore, individuals with this mitochondrial haplotype are in a high risk group. Because mitochondrial haplogroup U is found in 9.35% of the white United States population, there are more than 20 million individuals in this high risk group.
Link
January 12, 2006
Y chromosomes and Irish surnames
Human Genetics (online first)
Y-chromosomes and the extent of patrilineal ancestry in Irish surnames
Brian McEvoy and Daniel G. Bradley
Abstract Ireland has one of the oldest systems of patrilineal hereditary surnames in the world. Using the paternal co-inheritance of Y-chromosome DNA and Irish surnames, we examined the extent to which modern surname groups share a common male-line ancestor and the general applicability of Y-chromosomes in uncovering surname origins and histories. DNA samples were collected from 1,125 men, bearing 43 different surnames, and each was genotyped for 17 Y-chromosome short tandem repeat (STR) loci. A highly significant proportion of the observed Y-chromosome diversity was found between surnames demonstrating their demarcation of real and recent patrilineal kinship. On average, a man has a 30-fold increased chance of sharing a 17 STR Y-chromosome haplotype with another man of the same surname but the extent of congruence between the surname and haplotype varies widely between surnames and we attributed this to differences in the number of early founders. Some surnames such as O’Sullivan and Ryan have a single major ancestor, whereas others like Murphy and Kelly have numerous founders probably explaining their high frequency today. Notwithstanding differences in their early origins, all surnames have been extensively affected by later male introgession. None examined showed more than about half of current bearers still descended from one original founder indicating dynamic and continuously evolving kinship groupings. Precisely because of this otherwise cryptic complexity there is a substantial role for the Y-chromosome and a molecular genealogical approach to complement and expand existing sources.
Link
The matrilineal ancestry of Ashkenazi Jews
Update
You can read more about the findings in this CNN news story. You can also read the paper via Family Tree DNA (pdf).
The lead author, Doron Behar is associated with Family Tree DNA, so expect a "Jewish Daughters" type of test to become commercially available very soon, followed no doubt by wild speculation about the identity of the four mothers of Ashkenazi Jewry and their relationships with Biblical personalities.
Am J Hum Genet (online early)
The Matrilineal Ancestry of Ashkenazi Jewry: Portrait of a Recent Founder Event
Doron M. Behar et al.
Both the extent and location of the maternal ancestral deme from which the Ashkenazi Jewry arose remain obscure. Here, using complete sequences of the maternally inherited mitochondrial DNA (mtDNA), we show that close to one-half of Ashkenazi Jews, estimated at 8,000,000 people, can be traced back to only 4 women carrying distinct mtDNAs that are virtually absent in other populations, with the important exception of low frequencies among non-Ashkenazi Jews. We conclude that four founding mtDNAs, likely of Near Eastern ancestry, underwent major expansion(s) in Europe within the past millennium.
Link
New SNPs in haplogroup I
There are also four new markers within hg I, which together with a marker launched previously (S23) bring considerably improved resolution to the group as a whole. S31 is a new marker which unites haplogroups I1b and I1c and two I1* groups. S23, S30, S32 and S33 unite one of these I1* groups and hg I1c. Renaming of the groups will thus be required - I1b will become I1b1, one of the I1* groups becomes I1b*, the other I1* group becomes I1b2* and I1c will become I1b2a if the nomenclature is followed."The new I1b-S31 is a new link between Balkan and NW European haplogroup I chromosomes. Together with the previous announcement of haplogroup IJ-S22, the new SNP is helping us better trace the movements of the multiple waves of migrants into Europe through the Balkans.
January 11, 2006
New India Y-chromosome paper
India Acquired Language, not Genes, From WestBased on my readings so far, I believe that the Proto-Indo-Iranians and hence the ancestors of the first speakers of Indic speakers belonged primarily to haplogroups J2a and R1a1. Today, J2a is found approximately 4% of Hindus, and primarily in the upper castes where the signal of the founders of the caste system would be most evident.
Most modern Indians descended from South Asians, not invading Central Asian steppe dwellers, a new genetic study reports. The Indian subcontinent may have acquired agricultural techniques and languages—but it absorbed few genes—from the west, said Vijendra Kashyap, director of India's National Institute of Biologicals in Noida.
R1a1 is found across the caste hierarchy and in tribals and is more diverse in tribals and lower castes than in the upper castes. I suspect though, that the lack of informative subclades of R1a1 may mask multiple origins. Most of it is doubtlessly pre-Indo-European and probably pre-Neolithic, but some yet-undetected clade will be discovered that is of more recent origin. This will reflect the genetic contribution of Eurasian steppe groups which were themselves Indo-Europeanized at an earlier time from J2a-bearing populations that spread from Anatolia through the Balkans into Eastern Europe.
So, Hindus are indeed primarily descended from Pre-Indo-European populations, but they may still possess the signal of the arrival of the first Indic speakers. This will become even clearer once informative SNPs are discovered in haplogroup R1a1.
More to follow once I get a copy of the paper.
January 10, 2006
Excavation on Keros aims to solve riddle of Cycladic figurines

From CBSNews (excerpt):
"What is particularly impressive is not just the bulk of the finds, which is larger than the total from the rest of the Cyclades, but also that they were intentionally broken during ancient times," Sotirakopoulou said. "Therefore, this is a very important, a unique site."
The Cycladic culture _ a network of small, sometimes fortified farming and fishing settlements that traded with mainland Greece, Crete and Asia Minor _ is best known for its elegant artwork: mostly naked, elongated figures with their arms folded under their chest. The seafaring civilization was eclipsed in the second millennium B.C. by Crete and Mycenaean Greece.
...
Evidence from excavations in the '60s and 1980s failed to explain why the barren islet was so much more important in the 3rd millennium B.C. than its bigger, more hospitable neighbors.
"The prevailing explanation is that this was a sacred repository, a sort of pan-Cycladic sanctuary where people left objects within the framework of rituals which included their intentional smashing," said Sotirakopoulou.
She will participate in the summer's excavation together with Cambridge University professor Colin Renfrew and other experts.
Early human population centers in coastal habitats (?)
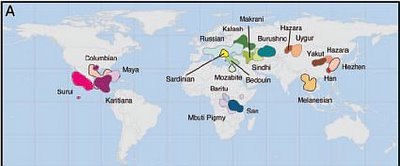 Examples of 50%-confidence kernels for population placement based on population average allele frequencies. Populations are: Columbian, Hezhen, Kalash, Karitiana, Lahu, Makrani, Mandega, Maya, Mbuti Pigmy, Melanesian, Mozabite, New Guinea, Orcadian, Palestinian, Pima, Russian, Sardinian, Uygur, Yakut, Yizu, and San. In most cases, a single kernel results, although, in populations Hezhen, Karitiana, and Pima, the kernel has broken into two or three regions.
Examples of 50%-confidence kernels for population placement based on population average allele frequencies. Populations are: Columbian, Hezhen, Kalash, Karitiana, Lahu, Makrani, Mandega, Maya, Mbuti Pigmy, Melanesian, Mozabite, New Guinea, Orcadian, Palestinian, Pima, Russian, Sardinian, Uygur, Yakut, Yizu, and San. In most cases, a single kernel results, although, in populations Hezhen, Karitiana, and Pima, the kernel has broken into two or three regions.Proc. Natl. Acad. Sci. USA, 10.1073/pnas.0507991103
Global genetic positioning: Evidence for early human population centers in coastal habitats
William Amos and Andrea Manica
For an alternative perspective on relationships among human populations, we combined genetic and geographic information, using allele frequency gradients to place populations and individuals on the globe. Reanalyzing published data on 51 worldwide populations [Rosenberg, N. A., Pritchard, J. K., Weber, J. L., Cann, H. M., Kidd, K. K., Zhivitovsky, L. A. & Feldman, M. W. (2002) Science 298, 2381-2385] reveals five geographic clusters lying in plausible sites either of early agricultural innovation or on ancient migration routes. Also, the inferred sites show significant association with coastlines, suggesting that most early humans lived near large bodies of water. Our approach is flexible, and developments should prove useful both for exploring historical demography and for the identification of likely origin for unknown forensic samples.
Link
January 08, 2006
News from the J world
Also, a new J2 Y-DNA project was started. Unfortunately, project haplotypes are not listed, but you can at least get a basic introduction into haplogroup J2, with more statistical analyses to come. The site contains a nice graphic of the phylogenetic correspondences of the new and old nomenclatures, which seems to be on the whole accurate, although the left-most paragroup should be labeled J2*(xDYS413<=18,J2h) (9 Jan: fixed).
The M410 project hosts the very informative Y haplogroup J database (yJdb) that seems to contain nearly all major published haplotypes of haplogroup J, a very convenient resource for cataloguing and searching J haplotypes. I will add yJdb to my list of genetic databases on the right sidebar.
Both projects seem to support haplotype submissions from regular people. An interesting feature of the the yJdb system is that it allows users to enter SNP results, unlike the mainstram ysearch and ybase databases. This is quite useful when trying to compare haplogroup names across different nomenclature systems and with different sets of genotyped markers.
If you are aware of other worthwhile efforts, feel free to leave a comment.
January 06, 2006
Y chromosomes of Golla subcastes

American Journal of Physical Anthropology (Early view)
Genetic diversity within a caste population of India as measured by Y-chromosome haplogroups and haplotypes: Subcastes of the Golla of Andhra Pradesh
R. John Mitchell et al.
Abstract
The extent of population subdivision based on 15 Y-chromosome polymorphisms was studied in seven subcastes of the Golla (Karnam, Pokanati, Erra, Doddi, Punugu, Puja, and Kurava), who inhabit the Chittoor district of southern Andhra Pradesh, India. These Golla subcastes are traditionally pastoralists, culturally homogeneous and endogamous. DNA samples from 146 Golla males were scored for seven unique event polymorphisms (UEPs) and eight microsatellites, permitting allocation of each into haplogroups and haplotypes, respectively. Genetic diversity (D) was high (range, 0.9048-0.9921), and most of the genetic variance (>91%) was explained by intrapopulation differences. Median-joining network analysis of microsatellite haplotypes demonstrated an absence of any structure according to subcaste affiliation. Superimposition of UEPs on this phylogeny, however, did create some distinct clusters, indicating congruence between haplotype and haplogroup phylogenies. Our results suggest many male ancestors for the Golla as well as for each of the subcastes. Genetic distances among the seven subcastes, based on autosomal markers (short tandem repeats and human leukocyte antigens) as well as those on the chromosome Y, indicate that the Kurava may not be a true subcaste of the Golla. Although this finding is based on a very small Kurava sample, it is in accordance with ethnohistorical accounts related by community elders. The Punugu was the first to hive off the main Golla group, and the most recently separated subcastes (Karnam, Erra, Doddi, and Pokanati) fissioned from the Puja. This phylogeny receives support from the analysis of autosomal microsatellites as well as HLA loci in the same samples. In particular, there is a significant correlation (r = 0.8569; P = 0.0097) between Y-chromosome- and autosomal STR-based distances.
Link
New reconstruction of Krapina 5 Neandertal
American Journal of Physical Anthropology (Early view)
New reconstruction of Krapina 5, a male Neandertal cranial vault from Krapina, Croatia
Rachel Caspari, Jakov Radovi
Abstract
The Neandertals from Krapina, Croatia represent some of the geologically oldest Neandertals known, and they comprise the largest Neandertal collection from a single site in the world. However, comparisons of the Krapina material with other, later Neandertals have been limited both because of their fragmentary condition and because the sample has a disproportionate number of females and/or young individuals. This paper presents a preliminary description of our new reconstruction of Krapina 5, an adult male, and provides comparisons with females from Krapina and with later Neandertal males from Western Europe. Like other hominid sites with large samples, there is considerable cranial variation at Krapina; we believe that some, but clearly not all of it is due to sexual dimorphism. Although Krapina 5 differs from the later males in a number of features, such as cranial thickness, cranial height, and sagittal curvature, it fits well within the male Neandertal range for most other metric variables, including cranial capacity.
Link
Mozart's skull and the DNA of the famous
So, will Wolfgang Amadeus join the ranks of the famous past and present whose patrilineal or matrilineal haplogroup is recorded? So far we have Genghis Khan [Y: C*(xC3c)], Somerled [Y: R1a1], Thomas Jefferson [Y: K2], the Manchu dynast Nurhachi [Y: C3c] (*), the founder of the Uí Néill dynasty [Y: R1b], the Tyrolean Ice Man [K], Cheddar Man [mt: U5a], Tsar Nicolas [mt: T], Spencer Wells [Y: R1b], and of course none others than Matt Lauer [Y: J2], Katie Couric [mt: K], Ann Curry [mt: N9a], and Al Roker [Y: E*(xE3a) ?] of the Today Show.
Update
Well the "clear result" that the scientists advertised is actually this:
The scientists said on Austrian television Sunday that the skeletons do not match the skull, and that the skeletons are also unrelated - creating a whole new mystery of who is buried the Mozart family crypt.(*) Whose fictional remains make a cameo in Indiana Jones and the Temple of Doom for those interested in movie trivia.
Update
See also Famous DNA.
Update
More here.
“I am quite disappointed that the mystery continues,” said Parsons. “All the samples from the three who were believed to be relatives of Mozart all had different mitochondrial DNA from each other, and from the Mozart skull. So if any one of them is an actual maternal relative of Mozart, it means that the skull is not Mozart’s. We don’t know if that is the case so the final analysis is inconclusive.
“We have attained definitive results from the skull,” said Parsons. “In the future, if anyone comes forward with an authentic matrilineal relative or a paternal relative, we now have ‘y’ chromosomal data and we will be in a position to make a confirmation. It’s considered to be known where Mozart’s sister Nannerl is buried, but I don’t know if there are any plans in Austria to act on that information and work another archeological exhumation.”
After several months of testing, the true identity of the skull remains inconclusive to be that of world-renowned 18th-century classical musical composer Mozart.
January 05, 2006
New dates for Vindija Neandertals: around 32-33kya
Revised direct radiocarbon dating of the Vindija G1 Upper Paleolithic Neandertals
Tom Higham et al.
The 1998/1999 direct dating of two Neandertal specimens from level G1 of Vindija Cave in Croatia to approximately 28,000 and approximately 29,000 radiocarbon (14C) years ago has led to interpretations concerning the late survival of Neandertals in south-central Europe, patterns of interaction between Neandertals and in-dispersing early modern humans in Europe, and complex biocultural scenarios for the earlier phases of the Upper Paleolithic. Given improvements, particularly in sample pretreatment techniques for bone radiocarbon samples, especially ultrafiltration of collagen samples, these Vindija G1 Neandertal fossils are redated to approximately 32,000-33,000 14C years ago and possibly earlier. These results and the recent redating of a number of purportedly old modern human skeletal remains in Europe to younger time periods highlight the importance of fine chronological control when studying this biocultural time period and the tenuous nature of monolithic scenarios for the establishment of modern humans and earlier phases of the Upper Paleolithic in Europe.
Link




