I remember how in the early days of online discussions of anthropology a constant topic of contention was whether African variation was the result of admixture, some of it within Africa, some of it from Caucasoids, or whether it was the result of climatic adaptation manifested in gradual clines (as opposed to clusters corresponding to physical types).
Well, I won't dismiss the role of climate altogether, but it's hard to argue for it much anymore now that we know that the two big fish in the African ocean of human diversity were the spread of Niger-Congo languages (from the west), and of Caucasoids (from the east) over the last few thousands of years, with a healthy seasoning of minor admixtures before and after. Once again it seems that old-style anthropology was right and the more fashionable and trendy attempts to dismiss it as "typology", "imposition of European colonialism through science" and the like were wrong.
eLife 2016;5:e15266
Admixture into and within sub-Saharan Africa
George BJ Busby et al.
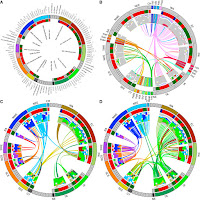
Similarity between two individuals in the combination of genetic markers along their chromosomes indicates shared ancestry and can be used to identify historical connections between different population groups due to admixture. We use a genome-wide, haplotype-based, analysis to characterise the structure of genetic diversity and gene-flow in a collection of 48 sub-Saharan African groups. We show that coastal populations experienced an influx of Eurasian haplotypes over the last 7000 years, and that Eastern and Southern Niger-Congo speaking groups share ancestry with Central West Africans as a result of recent population expansions. In fact, most sub-Saharan populations share ancestry with groups from outside of their current geographic region as a result of gene-flow within the last 4000 years. Our in-depth analysis provides insight into haplotype sharing across different ethno-linguistic groups and the recent movement of alleles into new environments, both of which are relevant to studies of genetic epidemiology.
Link
Showing posts with label ChromoPainter. Show all posts
Showing posts with label ChromoPainter. Show all posts
June 21, 2016
September 19, 2015
Recent admixture in contemporary West Eurasians
After applying Globetrotter to the world and to the British, a new study in Current Biology applies to the intermediately-sized region of West Eurasia. This is an open-access article, so go ahead and read it.
Current Biology DOI: http://dx.doi.org/10.1016/j.cub.2015.08.007
The Role of Recent Admixture in Forming the Contemporary West Eurasian Genomic Landscape
George B.J. Busby et al.
Over the past few years, studies of DNA isolated from human fossils and archaeological remains have generated considerable novel insight into the history of our species. Several landmark papers have described the genomes of ancient humans across West Eurasia, demonstrating the presence of large-scale, dynamic population movements over the last 10,000 years, such that ancestry across present-day populations is likely to be a mixture of several ancient groups [ 1–7 ]. While these efforts are bringing the details of West Eurasian prehistory into increasing focus, studies aimed at understanding the processes behind the generation of the current West Eurasian genetic landscape have been limited by the number of populations sampled or have been either too regional or global in their outlook [ 8–11 ]. Here, using recently described haplotype-based techniques [ 11 ], we present the results of a systematic survey of recent admixture history across Western Eurasia and show that admixture is a universal property across almost all groups. Admixture in all regions except North Western Europe involved the influx of genetic material from outside of West Eurasia, which we date to specific time periods. Within Northern, Western, and Central Europe, admixture tended to occur between local groups during the period 300 to 1200 CE. Comparisons of the genetic profiles of West Eurasians before and after admixture show that population movements within the last 1,500 years are likely to have maintained differentiation among groups. Our analysis provides a timeline of the gene flow events that have generated the contemporary genetic landscape of West Eurasia.
Link
March 18, 2015
British origins (Leslie et al. 2015)
The long-awaited paper on the People of the British Isles has just appeared in Nature. I will update this entry with more information.
UPDATE:
The authors write:
Most of the clusters are geographical, but some span different regions (e.g., the "yellow circle" cluster). The elephant in the room is the "red square" cluster which spans Central/South England. The authors write:
The authors draw conclusions on several historical episodes of British history. The big one is the extent of Anglo-Saxon ancestry:
Nature 519, 309–314 (19 March 2015) doi:10.1038/nature14230
The fine-scale genetic structure of the British population
Stephen Leslie et al.
Fine-scale genetic variation between human populations is interesting as a signature of historical demographic events and because of its potential for confounding disease studies. We use haplotype-based statistical methods to analyse genome-wide single nucleotide polymorphism (SNP) data from a carefully chosen geographically diverse sample of 2,039 individuals from the United Kingdom. This reveals a rich and detailed pattern of genetic differentiation with remarkable concordance between genetic clusters and geography. The regional genetic differentiation and differing patterns of shared ancestry with 6,209 individuals from across Europe carry clear signals of historical demographic events. We estimate the genetic contribution to southeastern England from Anglo-Saxon migrations to be under half, and identify the regions not carrying genetic material from these migrations. We suggest significant pre-Roman but post-Mesolithic movement into southeastern England from continental Europe, and show that in non-Saxon parts of the United Kingdom, there exist genetically differentiated subgroups rather than a general ‘Celtic’ population.
Link
UPDATE:
The authors write:
Consistent with earlier studies of the UK, population structure within the PoBI collection is very limited. The average of the pairwise FST estimates between each of the 30 sample collection districts is 0.0007, with a maximum of 0.003 (Supplementary Table 1).These are extremely small differences in the European (let alone global) context. So, the British are, overall, a very homogeneous population. This is what led the researchers to use methods such as ChromoPainter/ fineStructure/ Globetrotter that can squeeze out fine-scale population structure by exploiting linkage disequilibrium. Thus, the authors are able to detect 17 main clusters of the British.
Most of the clusters are geographical, but some span different regions (e.g., the "yellow circle" cluster). The elephant in the room is the "red square" cluster which spans Central/South England. The authors write:
There is a single large cluster (red squares) that covers most of central and southern England and extends up the east coast. Notably, even at the finest level of differentiation returned by fineSTRUCTURE (53 clusters), this cluster remains largely intact and contains almost half the individuals (1,006) in our study.The authors then tried to infer the ancestry of the British clusters in terms of continental European clusters, which is to be published separately. In the plot on the right, you see the British clusters (columns) and their continental European sources (rows). The authors observe that clusters that are widely represented in Britain are likely to be older, while those that are missing in some populations are likely to be younger, because they didn't have the chance to spread across Britain. For example, a couple of Norwegian clusters are strongly represented in the Orkney islands, and these are likely to reflect Viking colonization.
The authors draw conclusions on several historical episodes of British history. The big one is the extent of Anglo-Saxon ancestry:
After the Saxon migrations, the language, place names, cereal crops and pottery styles all changed from that of the existing (Romano-British) population to those of the Saxon migrants. There has been ongoing historical and archaeological controversy about the extent to which the Saxons replaced the existing Romano-British populations. Earlier genetic analyses, based on limited samples and specific loci, gave conflicting results. With genome-wide data we can resolve this debate. Two separate analyses (ancestry profiles and GLOBETROTTER) show clear evidence in modern England of the Saxon migration, but each limits the proportion of Saxon ancestry, clearly excluding the possibility of long-term Saxon replacement. We estimate the proportion of Saxon ancestry in Cent./S England as very likely to be under 50%, and most likely in the range of 10–40%.Two other details are the lack of Danish Viking ancestry in England:
In particular, we see no clear genetic evidence of the Danish Viking occupation and control of a large part of England, either in separate UK clusters in that region, or in estimated ancestry profiles, suggesting a relatively limited input of DNA from the Danish Vikings and subsequent mixing with nearby regions, and clear evidence for only a minority Norse contribution (about 25%) to the current Orkney population.And, the absence of a unified pre-Saxon "Celtic" population. What seems to unify "Celts" is lower levels/absence of the Saxon influence, rather than belonging to a homogeneous "Celtic" population:
We saw no evidence of a general ‘Celtic’ population in non-Saxon parts of the UK. Instead there were many distinct genetic clusters in these regions, some amongst the most different in our study, in the sense of being most separated in the hierarchical clustering tree in Fig. 1. Further, the ancestry profile of Cornwall (perhaps expected to resemble other Celtic clusters) is quite different from that of the Welsh clusters, and much closer to that of Devon, and Cent./S England. However, the data do suggest that the Welsh clusters represent populations that are more similar to the early post-Ice-Age settlers of Britain than those from elsewhere in the UK.Unfortunately, the authors have decided not to make their data publicly available. This is very unfortunate, and will keep this research out of the hands of many people who would be interested in it and who would be interested in analyzing this data. I can already guess the disappointment of people of British ancestry from around the world who have a genealogical interest in tracing their British ancestors to particular areas of the UK. Apparently, the data is deposited in the EGA archive, access requires red tape, and is apparently limited to institutional researchers. Thus, this data, perhaps the richest genetic survey of any country to date, will not be fully utilized to further science.
Nature 519, 309–314 (19 March 2015) doi:10.1038/nature14230
The fine-scale genetic structure of the British population
Stephen Leslie et al.
Fine-scale genetic variation between human populations is interesting as a signature of historical demographic events and because of its potential for confounding disease studies. We use haplotype-based statistical methods to analyse genome-wide single nucleotide polymorphism (SNP) data from a carefully chosen geographically diverse sample of 2,039 individuals from the United Kingdom. This reveals a rich and detailed pattern of genetic differentiation with remarkable concordance between genetic clusters and geography. The regional genetic differentiation and differing patterns of shared ancestry with 6,209 individuals from across Europe carry clear signals of historical demographic events. We estimate the genetic contribution to southeastern England from Anglo-Saxon migrations to be under half, and identify the regions not carrying genetic material from these migrations. We suggest significant pre-Roman but post-Mesolithic movement into southeastern England from continental Europe, and show that in non-Saxon parts of the United Kingdom, there exist genetically differentiated subgroups rather than a general ‘Celtic’ population.
Link
February 13, 2014
Human admixture common in human history (Hellenthal et al. 2014)
A string of recent papers argued for admixture in human populations at time scales from the Middle Pleistocene to recent centuries. A new paper in Science makes the point convincingly for extensive admixture in humans over the last few thousand years. The authors include the creators of Chromopainter/fineStructure software; the new "Globetrotter" method appears to be a natural extension of that method that seemed to work wonderfully well except for the limitation of producing only a tree of the studied populations.
The paper has a companion website in which you can look up the admixture history of individual populations.
While reading this study, it is important to remember its limitations. Two are immediately obvious: (i) admixture events can only be detected for the last few thousand years, as this method depends on pattern of linkage disequilibrium which decays exponentially with time due to recombination, and (ii) detection of admixture seems to depend on the presence of maximally differentiated populations from the edges of the human geographical range; for example, the Japanese appear unadmixed even though they are clearly of dual Jomon/Yayoi ancestry. On the other hand, the method does detect the admixture present in the San at a similar time scale.
The case of Northwestern Europe appears especially striking as none of the populations from the region show evidence of admixture. This may be because the mixtures taking place there (e.g., between "Celts" and "Anglo-Saxons" in Great Britain) involved populations that were not strongly differentiated. Alternatively, population admixture history may have preceded the last few thousand years and is thus beyond the temporal scope of this method.
An exception to the rule that populations at the edges of the human range appear to be unadmixed are the Armenians who appear to be the only * between the Atlantic and Pacific in Figure 2D (shown at the beginning of this post). The companion site lists their status as "uncertain".
Other results are more questionable; for example, the authors assert that Sardinians are an admixed population with one side being "Egyptian-like" and the other "French-like" whereas the ancient DNA evidence as it stands would rather indicate that Sardinians are the best approximation of Neolithic Europeans currently in existence and so are more likely to (mostly) possess a gene pool that traces back to ~8-9 thousand years in Europe. It will be quite the surprise if so many Europeans from 5kya or earlier look like modern Sardinians and ancient Sardinians don't!
The analysis of Eastern Europe is particularly interesting as it documents three way admixture (Northern/Southern/NE Asian) in most populations but two way admixture (Northern/Southern) in Greeks, estimated at ~37%. The authors claim that this is related to the Slavs, which seems reasonable given the 1,054AD age estimate. On the other hand, according to the companion website, the southern element in Greeks is inferred to be Cypriot-like and it's far from clear that the pre-Slavic population of Greece was Cypriot-like or indeed represented by any of the populations in the authors' dataset.
The three-way admixture in much of eastern Europe is not particularly surprising as history furnishes ample evidence for groups of steppe origin in the region during historical times. Some bequeathed their both language and name (e.g., Magyars), others only their name (e.g., Bulgarians) on the local Europeans, but records indicate a widespread presence of "eastern" groups in Europe from the time of the Huns to that of the Ottomans. A study of late Antique eastern Europeans from the Baltic to the Aegean may help better document how the twin phenomena of the eastern invasions and the spread of the Slavs shaped the present-day genetic diversity of the region.
I suspect that a few ancient samples will be far more informative for understanding the recent history of our species than the most sophisticated modeling of modern populations. Nonetheless, it's great to have a new method that maximizes what can be learned about the past from the messy palimpsest of the present.
Science 14 February 2014: Vol. 343 no. 6172 pp. 747-751 DOI: 10.1126/science.1243518
A Genetic Atlas of Human Admixture History
Garrett Hellenthal et al.
Modern genetic data combined with appropriate statistical methods have the potential to contribute substantially to our understanding of human history. We have developed an approach that exploits the genomic structure of admixed populations to date and characterize historical mixture events at fine scales. We used this to produce an atlas of worldwide human admixture history, constructed by using genetic data alone and encompassing over 100 events occurring over the past 4000 years. We identified events whose dates and participants suggest they describe genetic impacts of the Mongol empire, Arab slave trade, Bantu expansion, first millennium CE migrations in Eastern Europe, and European colonialism, as well as unrecorded events, revealing admixture to be an almost universal force shaping human populations.
Link
The paper has a companion website in which you can look up the admixture history of individual populations.
While reading this study, it is important to remember its limitations. Two are immediately obvious: (i) admixture events can only be detected for the last few thousand years, as this method depends on pattern of linkage disequilibrium which decays exponentially with time due to recombination, and (ii) detection of admixture seems to depend on the presence of maximally differentiated populations from the edges of the human geographical range; for example, the Japanese appear unadmixed even though they are clearly of dual Jomon/Yayoi ancestry. On the other hand, the method does detect the admixture present in the San at a similar time scale.
The case of Northwestern Europe appears especially striking as none of the populations from the region show evidence of admixture. This may be because the mixtures taking place there (e.g., between "Celts" and "Anglo-Saxons" in Great Britain) involved populations that were not strongly differentiated. Alternatively, population admixture history may have preceded the last few thousand years and is thus beyond the temporal scope of this method.
An exception to the rule that populations at the edges of the human range appear to be unadmixed are the Armenians who appear to be the only * between the Atlantic and Pacific in Figure 2D (shown at the beginning of this post). The companion site lists their status as "uncertain".
Other results are more questionable; for example, the authors assert that Sardinians are an admixed population with one side being "Egyptian-like" and the other "French-like" whereas the ancient DNA evidence as it stands would rather indicate that Sardinians are the best approximation of Neolithic Europeans currently in existence and so are more likely to (mostly) possess a gene pool that traces back to ~8-9 thousand years in Europe. It will be quite the surprise if so many Europeans from 5kya or earlier look like modern Sardinians and ancient Sardinians don't!
The analysis of Eastern Europe is particularly interesting as it documents three way admixture (Northern/Southern/NE Asian) in most populations but two way admixture (Northern/Southern) in Greeks, estimated at ~37%. The authors claim that this is related to the Slavs, which seems reasonable given the 1,054AD age estimate. On the other hand, according to the companion website, the southern element in Greeks is inferred to be Cypriot-like and it's far from clear that the pre-Slavic population of Greece was Cypriot-like or indeed represented by any of the populations in the authors' dataset.
The three-way admixture in much of eastern Europe is not particularly surprising as history furnishes ample evidence for groups of steppe origin in the region during historical times. Some bequeathed their both language and name (e.g., Magyars), others only their name (e.g., Bulgarians) on the local Europeans, but records indicate a widespread presence of "eastern" groups in Europe from the time of the Huns to that of the Ottomans. A study of late Antique eastern Europeans from the Baltic to the Aegean may help better document how the twin phenomena of the eastern invasions and the spread of the Slavs shaped the present-day genetic diversity of the region.
I suspect that a few ancient samples will be far more informative for understanding the recent history of our species than the most sophisticated modeling of modern populations. Nonetheless, it's great to have a new method that maximizes what can be learned about the past from the messy palimpsest of the present.
Science 14 February 2014: Vol. 343 no. 6172 pp. 747-751 DOI: 10.1126/science.1243518
A Genetic Atlas of Human Admixture History
Garrett Hellenthal et al.
Modern genetic data combined with appropriate statistical methods have the potential to contribute substantially to our understanding of human history. We have developed an approach that exploits the genomic structure of admixed populations to date and characterize historical mixture events at fine scales. We used this to produce an atlas of worldwide human admixture history, constructed by using genetic data alone and encompassing over 100 events occurring over the past 4000 years. We identified events whose dates and participants suggest they describe genetic impacts of the Mongol empire, Arab slave trade, Bantu expansion, first millennium CE migrations in Eastern Europe, and European colonialism, as well as unrecorded events, revealing admixture to be an almost universal force shaping human populations.
Link
June 06, 2013
What is a population?
A nice overview paper comparing various methods used in population genetics has appeared on the arXiv.
arXiv:1306.0701 [q-bio.PE]
Populations in statistical genetic modelling and inference
Daniel John Lawson
What is a population? This review considers how a population may be defined in terms of understanding the structure of the underlying genetics of the individuals involved. The main approach is to consider statistically identifiable groups of randomly mating individuals, which is well defined in theory for any type of (sexual) organism. We discuss generative models using drift, admixture and spatial structure, and the ancestral recombination graph. These are contrasted with statistical models for inference, principle component analysis and other `non-parametric' methods. The relationships between these approaches are explored with both simulated and real-data examples. The state-of-the-art practical software tools are discussed and contrasted. We conclude that populations are a useful theoretical construct that can be well defined in theory and often approximately exist in practice.
Link
arXiv:1306.0701 [q-bio.PE]
Populations in statistical genetic modelling and inference
Daniel John Lawson
What is a population? This review considers how a population may be defined in terms of understanding the structure of the underlying genetics of the individuals involved. The main approach is to consider statistically identifiable groups of randomly mating individuals, which is well defined in theory for any type of (sexual) organism. We discuss generative models using drift, admixture and spatial structure, and the ancestral recombination graph. These are contrasted with statistical models for inference, principle component analysis and other `non-parametric' methods. The relationships between these approaches are explored with both simulated and real-data examples. The state-of-the-art practical software tools are discussed and contrasted. We conclude that populations are a useful theoretical construct that can be well defined in theory and often approximately exist in practice.
Link
August 01, 2012
Let's play ASHG 2012 title imputation! (+open science miscellanea)
It's that time of year again, and the titles for the ASHG 2012 presentations have just been posted online. Well, part of them anyway; the dreaded (...) have made another appearance. Still it is fun to try to guess what each contribution is about, and we'll only have to wait ~1 month for the abstract text.
In related news, Ewen Callaway reports on the trend (?) for biologists to put their unpublished work in arXiv. My own views are strictly for open science, so I applaud the people who are dragging their disciplines into the 21st century.
Finally, recent initiatives in the UK and the EU will mandate open access for work funded by research agencies. This is a good step in the right direction, but a very incomplete one: open access solves the problem of ensuring wide dissemination of new science, but merely shifts the flow of public money rather than sever it. With open access Government->University Library->Journal is replaced by Government->Research Agency->Scholar->Journal.
Moreover, open access does not address the more fundamental issue of how journals impede scientific progress by imposing the antiquated pre-publication peer review process. The sky hasn't fallen over the heads of physicists who post their work on arXiv when they're done with it and carry out post-arXiv publication peer review. So, it probably won't fall on the heads of biologists who do the same either.
A good example of this is the recent work on ChromoPainter/fineSTRUCTURE that appeared months before publication: lots of people -including myself- started using their software right away, which spurred new insight, and they got their peer-reviewed publication too. More recently, a group of independent researchers co-ordinated their efforts in public to hack 1000 Genomes data, discovered and validated new SNPs, and they got their publication too. Open science works, so everyone should try it!
In related news, Ewen Callaway reports on the trend (?) for biologists to put their unpublished work in arXiv. My own views are strictly for open science, so I applaud the people who are dragging their disciplines into the 21st century.
Finally, recent initiatives in the UK and the EU will mandate open access for work funded by research agencies. This is a good step in the right direction, but a very incomplete one: open access solves the problem of ensuring wide dissemination of new science, but merely shifts the flow of public money rather than sever it. With open access Government->University Library->Journal is replaced by Government->Research Agency->Scholar->Journal.
Moreover, open access does not address the more fundamental issue of how journals impede scientific progress by imposing the antiquated pre-publication peer review process. The sky hasn't fallen over the heads of physicists who post their work on arXiv when they're done with it and carry out post-arXiv publication peer review. So, it probably won't fall on the heads of biologists who do the same either.
A good example of this is the recent work on ChromoPainter/fineSTRUCTURE that appeared months before publication: lots of people -including myself- started using their software right away, which spurred new insight, and they got their peer-reviewed publication too. More recently, a group of independent researchers co-ordinated their efforts in public to hack 1000 Genomes data, discovered and validated new SNPs, and they got their publication too. Open science works, so everyone should try it!
March 26, 2012
Similarity matrices and clustering (Lawson and Falush)
Lawson and Falush have a new review paper on different clustering methods using haplotype data such as their own ChromoPainter/fineSTRUCTURE methodology, as well as the MCLUST/fastIBD methods that I started playing with a while back.
I won't have much time for the next few days to comprehensively review this new work, but I will add one data point to the discussion, by pointing to my ChromoPainter and fastIBD analyses over the same dataset. I will also add any further comments on this blog post, once I get the opportunity to read the paper.
Another point that needs to be made is how commendable the ChromoPainter folks' attitude towards the topic has been. Not only did they post their ChromoPainter preprint and software online months before their original paper was published, but they quickly jumped on my comments and suggestions on their paper to write their new review paper, making at available as a preprint itself. I'm guessing this saved about a year or two over what would have been possible if all the formalities of "traditional" publishing had been observed. It's also a very nice example of synergy between professional and amateur science, that the Internet and social media has made possible.
Similarity matrices and clustering algorithms for population identification using genetic data
Daniel John Lawson and Daniel Falush
Abstract
A large number of algorithms have been developed to identify population
structure from genetic data. Recent results show that the information used
by both model-based clustering methods and Principal Components Analysis
can be summarised by a matrix of pairwise similarity measures between
individuals. Similarity matrices have been constructed in a number of ways,
usually treating markers as independent but differing in the weighting given
to polymorphisms of different frequencies. Additionally, methods are now being
developed that better exploit the power of genome data by taking linkage
into account. We review several such matrices and evaluate their ‘information
content’. A two-stage approach for population identification is to first construct
a similarity matrix, and then perform clustering. We review a range
of common clustering algorithms, and evaluate their performance through a
simulation study. The clustering step can be performed either directly, or
after using a dimension reduction technique such as Principal Components
Analysis, which we find substantially improves the performance of most algorithms.
Based on these results, we describe the population structure signal
contained in each similarity matrix, finding that accounting for linkage leads
to significant improvements for sequence data. We also perform a comparison
on real data, where we find that population genetics models outperform
generic clustering approaches, particularly in regards to robustness against
features such as relatedness between individuals.
Link
I won't have much time for the next few days to comprehensively review this new work, but I will add one data point to the discussion, by pointing to my ChromoPainter and fastIBD analyses over the same dataset. I will also add any further comments on this blog post, once I get the opportunity to read the paper.
Another point that needs to be made is how commendable the ChromoPainter folks' attitude towards the topic has been. Not only did they post their ChromoPainter preprint and software online months before their original paper was published, but they quickly jumped on my comments and suggestions on their paper to write their new review paper, making at available as a preprint itself. I'm guessing this saved about a year or two over what would have been possible if all the formalities of "traditional" publishing had been observed. It's also a very nice example of synergy between professional and amateur science, that the Internet and social media has made possible.
Similarity matrices and clustering algorithms for population identification using genetic data
Daniel John Lawson and Daniel Falush
Abstract
A large number of algorithms have been developed to identify population
structure from genetic data. Recent results show that the information used
by both model-based clustering methods and Principal Components Analysis
can be summarised by a matrix of pairwise similarity measures between
individuals. Similarity matrices have been constructed in a number of ways,
usually treating markers as independent but differing in the weighting given
to polymorphisms of different frequencies. Additionally, methods are now being
developed that better exploit the power of genome data by taking linkage
into account. We review several such matrices and evaluate their ‘information
content’. A two-stage approach for population identification is to first construct
a similarity matrix, and then perform clustering. We review a range
of common clustering algorithms, and evaluate their performance through a
simulation study. The clustering step can be performed either directly, or
after using a dimension reduction technique such as Principal Components
Analysis, which we find substantially improves the performance of most algorithms.
Based on these results, we describe the population structure signal
contained in each similarity matrix, finding that accounting for linkage leads
to significant improvements for sequence data. We also perform a comparison
on real data, where we find that population genetics models outperform
generic clustering approaches, particularly in regards to robustness against
features such as relatedness between individuals.
Link
February 22, 2012
ChromoPainter/fineSTRUCTURE analysis of select South Asian/West Eurasian populations
This is the final result of the analysis mentioned in this previous post on the Kalash, using all 22 chromosomes.
Due to the quadratic running time of ChromoPainter, I took a random sample of 15 individuals from every included population with more than15 individuals. The final set included 392 individuals. It appears that a set of ~400 individuals/~260k SNPs can be processed in about 2 weeks on a single thread.
The raw chunkcounts between all individuals can be obtained from here.
The heatmap can be seen below:
The principal components analysis, shows the familiar West-to-South Asia cline:
More information can be found in the spreadsheet, including:
Finally, in the RAR file you can find some plots of Z scores (by row) for the different population.
For example, here is a list of donors for the Kalash population; the order is slightly different compared to the teaser, but the overall pattern is the same.
Compare with an outbred population, such as the Armenians:
Due to the quadratic running time of ChromoPainter, I took a random sample of 15 individuals from every included population with more than15 individuals. The final set included 392 individuals. It appears that a set of ~400 individuals/~260k SNPs can be processed in about 2 weeks on a single thread.
The raw chunkcounts between all individuals can be obtained from here.
The heatmap can be seen below:
The principal components analysis, shows the familiar West-to-South Asia cline:
More information can be found in the spreadsheet, including:
- How many individuals from each population were assigned to each of 51 clusters
- Individual assignments of all 392 individuals
- Raw chunkcounts between all 33 different populations
- Z scores of the above (by row)
- Z scores of the above (by column)
- by row: scan each line to see which populations (columns) are the bigger donors for each row.
- by column: scan each column to see which populations (rows) are the bigger recipients for each column.
Finally, in the RAR file you can find some plots of Z scores (by row) for the different population.
For example, here is a list of donors for the Kalash population; the order is slightly different compared to the teaser, but the overall pattern is the same.
February 18, 2012
A teaser on the Kalash
UPDATE (Feb 22): The complete analysis can be found here.
Razib has a post on Kalash on the human tree. As it happens, I am in the middle of a ChromoPainter/fineSTRUCTURE analysis of a broad dataset designed to explore certain mysteries that have often come up in my previous experiments. Barring the unexpected, the analysis should be completed sometime next week.
Below you can see the normalized number of "chunks" donated by various populations to the Kalash. First, we normalize including intra-Kalash sharing:
Notice the extreme intra-Kalash haplotype sharing: Kalash individuals are recipient of "chunks" from other Kalash individuals ~5 standard deviations more often than the mean over this set of populations.
However, if we igonore intra-Kalash haplotype sharing, then the donor populations are:
Of particular interest is the fact that all West Asian populations appear higher on the donor list than all Northern European ones, which confirms, using a haplotype-based approach, my previous inference that the Ancestral North Indian (ANI) component is related to West Asians.
Razib has a post on Kalash on the human tree. As it happens, I am in the middle of a ChromoPainter/fineSTRUCTURE analysis of a broad dataset designed to explore certain mysteries that have often come up in my previous experiments. Barring the unexpected, the analysis should be completed sometime next week.
Below you can see the normalized number of "chunks" donated by various populations to the Kalash. First, we normalize including intra-Kalash sharing:
Notice the extreme intra-Kalash haplotype sharing: Kalash individuals are recipient of "chunks" from other Kalash individuals ~5 standard deviations more often than the mean over this set of populations.
However, if we igonore intra-Kalash haplotype sharing, then the donor populations are:
Of particular interest is the fact that all West Asian populations appear higher on the donor list than all Northern European ones, which confirms, using a haplotype-based approach, my previous inference that the Ancestral North Indian (ANI) component is related to West Asians.
Subscribe to:
Comments (Atom)