July 31, 2005
Debunking the concept of 'race' ... not
Of course, these results actually prove the existence of race. It would be interesting to see how many of the students had anything other than the majority of their ancestry from their self-identified racial group; my guess, based on many autosomal studies to date, is that this percentage would be minimal.
It is of course expected that individuals will have variable proportions of admixture. After all, races are not species, and people from different races have interbed in the past and are continuing to interbreed in the present.
Genetic testing has in fact vindicated the findings of traditional physical anthropology. Physical anthropologists have long believed in continental races, although they may have differed as to the number of these races. Whenever individuals from around the world are clustered based on a large number of loci, invariably the major races emerge as clusters of genetic similarity. Whenever the self-reported race of individuals is compared with their "genotypic" cluster, invariably the two agree.
In regions of the world such as Central Asia, East Africa, or Latin America where traditional physical anthropology has believed that intermixture of races has created mixed-race populations, genetics has invariably shown the hybridity of these populations. Groups such as African Americans who were known to have acquired a large degree of Caucasoid admixture due to the social definition of a "black" in the United States, have been shown to be a group with around 20-25% European admixture.
So, the concept of 'race' has not been debunked by modern genetics. Rather, it has been victoriously confirmed.
Update:
John Hawks has much to say about this "debunking" of race, also addressing some problems of the DNA Print test.
July 28, 2005
Pre-Roman Iberian mtDNA
The most frequent haplogroup is H (52.9%), followed by U (17.6%), J (11.8%), and pre-HV, K and T at the same frequency (5.9%). No samples were found to correspond to other haplogroups that are widely present in the Iberian peninsula populations (Table 7), such as V, X, I or W. The North African U6 subhaplogroup and Sub-Saharan African L lineages are also absent from the ancient Iberians analyzed so far; therefore, the possible entry of U6 lineages prior to the Muslim conquest in the 8th century A.D., as suggested by some authors, remains unproven. However, it is recognized that the sample size is at present too small to exclude any competing hypothesis about a possible North African genetic contribution to the genesis of the Iberian peninsula populations.
This appears to be in general agreement with the recent paper on the origin of African mtDNA in Iberia. The absence of mtDNA haplogroup V, common today in Iberia, is confirmed. Interestingly haplogroup H1, which is a subclade of the frequent haplogroup H was also found in the prehistoric sample. H1 is believed to have originated in Iberia.
Annals of Human Genetics (Early View)
The Genetics of the Pre-Roman Iberian Peninsula: A mtDNA Study of Ancient Iberians
M. L. Sampietro et al.
Summary
The Iberians developed a surprisingly sophisticated culture in the Mediterranean coast of the Iberian Peninsula from the 6th century BC until their conquest by the Romans in the 2nd century BC. They spoke and wrote a non-Indo-European language that still cannot be understood; their origins and relationships with other non-Indo-European peoples, like the Etruscans, are unclear, since their funerary practices were based on the cremation of bodies, and therefore anthropology has been unable to approach the study of this people. We have retrieved mitochondrial DNA (mtDNA) from a few of the scarce skeletal remains that have been preserved, some of them belonging to ritualistically executed individuals. The most stringent authentication criteria proposed for ancient DNA, such as independent replication, amino-acid analysis, quantitation of template molecules, multiple extractions and cloning of PCR products, have been followed to obtain reliable sequences from the mtDNA hypervariable region 1 (HVR1), as well as some haplogroup diagnostic SNPs. Phylogeographic analyses show that the haplogroup composition of the ancient Iberians was very similar to that found in modern Iberian Peninsula populations, suggesting a long-term genetic continuity since pre-Roman times. Nonetheless, there is less genetic diversity in the ancient Iberians than is found among modern populations, a fact that could reflect the small population size at the origin of the population sampled, and the heterogenic tribal structure of the Iberian society. Moreover, the Iberians were not especially closely related to the Etruscans, which points to considerable genetic heterogeneity in Pre-Roman Western Europe.
Link
Diversity of tribal Indians
Diversity and Divergence Among the Tribal Populations of India
W.S. Watkins et al.
Summary
Tribal populations of the Indian subcontinent have been of longstanding interest to anthropologists and human geneticists. To investigate the relationship of Indian tribes to Indian castes and continental populations, we analyzed 45 unlinked autosomal STR loci in 9 tribal groups, 8 castes, and 18 populations from Africa, Europe and East Asia. South Indian tribal populations demonstrate low within-population heterozygosity (range: 0.54 - 0.69), while tribal populations sampled further north and east have higher heterozygosity (range: 0.69 - 0.74). Genetic distance estimates show that tribal Indians are more closely related to caste Indians than to other major groups. Between-tribe differentiation is high and exceeds that for eight sub-Saharan African populations (4.8% vs. 3.7%). Telugu-speaking populations are less differentiated than non-Telugu speakers (FST: 0.029 vs. 0.079), but geographic distance was not predictive of genetic affinity between tribes. South Indian tribes show significant population structure, and individuals can be clustered statistically into groups that correspond with their tribal affiliation. These results are consistent with high levels of genetic drift and isolation in Indian tribal populations, particularly those of South India, and they imply that these populations may be potential candidates for linkage disequilibrium and association mapping.
Link
Ancient European DNA mutation causes ataxia
Mitochondrial DNA Polymerase W748S Mutation: A Common Cause of Autosomal Recessive Ataxia with Ancient European Origin
Anna H. Hakonen et al.
Mutations in the catalytic subunit of the mitochondrial DNA polymerase γ (POLG) have been found to be an important cause of neurological disease. Recently, we and collaborators reported a new neurodegenerative disorder with autosomal recessive ataxia in four patients homozygous for two amino acid changes in POLG: W748S in cis with E1143G. Here, we studied the frequency of this allele and found it to be among the most common genetic causes of inherited ataxia in Finland. We identified 27 patients with mitochondrial recessive ataxia syndrome (MIRAS) from 15 Finnish families, with a carrier frequency in the general population of 1 : 125. Since the mutation pair W748S+E1143G has also been described in European patients, we examined the haplotypes of 13 non-Finnish, European patients with the W748S mutation. Haplotype analysis revealed that all the chromosomes carrying these two changes, in patients from Finland, Norway, the United Kingdom, and Belgium, originate from a common ancient founder. In Finland and Norway, long, common, northern haplotypes, outside the core haplotype, could be identified. Despite having identical homozygous mutations, the Finnish patients with this adult- or juvenile-onset disease had surprisingly heterogeneous phenotypes, albeit with a characteristic set of features, including ataxia, peripheral neuropathy, dysarthria, mild cognitive impairment, involuntary movements, psychiatric symptoms, and epileptic seizures. The high carrier frequency in Finland, the high number of patients in Norway, and the ancient European founder chromosome indicate that this newly identified ataxia should be considered in the first-line differential diagnosis of progressive ataxia syndromes.
Link
July 22, 2005
Reliability of P25 marker
Forensic Sci Int. 2005 Jul 15; [Epub ahead of print]
The case of the unreliable SNP: Recurrent back-mutation of Y-chromosomal marker P25 through gene conversion.
Adams SM et al.
The Y-chromosomal binary marker P25 is a paralogous sequence variant, rather than a SNP: three copies of the P25 sequence lie within the giant palindromic repeats on Yq, and one copy has undergone a C to A transversion to define haplogroup R1b (designated C/C/A). Since gene conversion is known to be active in the palindromic repeats, we reasoned that P25 might be liable to back-mutation by gene conversion, yielding the ancestral state C/C/C. Through analysis of a set of binary markers in Y-chromosomes in two large samples from Great Britain and the Iberian Peninsula we show that such conversion events have occurred at least twice, and provide preliminary evidence that the reverse conversion event (yielding C/A/A) has also occurred. Because of its inherent instability, we suggest that P25 be used with caution in forensic studies, and perhaps replaced with the more reliable binary marker M269.
Link
July 20, 2005
Structure of Mongoloid populations
Human Genetics (Online first)
Use of autosomal loci for clustering individuals and populations of East Asian origin
Jong-Jin Kim et al.
Abstract We studied the genetic relationships among East Asian populations based on allele frequency differences to clarify the relative similarities of East Asian populations with a specific focus on the relationships among the Koreans, the Japanese, and the Chinese populations known to be genetically similar. The goal is to find markers appropriate for differentiating among the specific populations. In this study, no prior data existed for Koreans and the markers were selected to differentiate Chinese and Japanese. We typed, using AB TaqMan assays, single-nucleotide polymorphisms (SNPs) at 43 highly selected mostly independent diallelic sites, on 386 individuals from eight East Asian populations (Han Chinese from San Francisco, Han Chinese from Taiwan, Hakka, Koreans, Japanese, Ami, Atayal, and Cambodians) and one Siberian population (Yakut). We inferred group membership of individuals using a model-based clustering method implemented by the STRUCTURE program and population clustering by using computer programs DISTANCE, NEIGHBOR, LSSEARCH, and DRAWTREE, respectively, calculating genetic distances among populations, calculating neighbor-joining and least-squares trees, and drawing the calculated trees. On average 52% of individuals in the three Chinese groups were assigned into one cluster, and, respectively, 78 and 69% of Koreans and Japanese into a different cluster. Koreans differentiated from the Chinese groups and clustered with the Japanese in the principal component analysis (PCA) and in the best least-squares tree. The majority of Koreans were difficult to distinguish from the Japanese. This study shows that a relatively few highly selected markers can, within limits, differentiate between closely related populations.
Link
Y-haplogroups of some European population
- Detection of 1 haplogroup A* and 2 BCD chromosomes in Germany (3.2% non-Caucasoid paternal admixture)
- Detection of 1 E*(xE3b) chromosome in Austria (0.8% non-Caucasoid paternal admixture)
- Detection of 1 R1a*(xR1a1) chromosome in Norway; usually R1a chromosomes also harbor the defining mutation for R1a1, but it appears that this is not always the case.
Forensic Science International (Article in Press, Corrected Proof)
A collaborative study of the EDNAP group regarding Y-chromosome binary polymorphism analysis
María Brion et al.
Abstract
A collaborative study was carried out by the European DNA Profiling Group (EDNAP) in order to evaluate the performance of Y-chromosome binary polymorphism analysis in different European laboratories. Four blood samples were sent to the laboratories, to be analysed for 11 Y-chromosome single nucleotide polymorphisms (SNPs): SRY-1532, M40, M35, M213, M9, 92R7, M17, P25, M18, M153 and M167. All the labs were also asked to submit a population study including these markers.
All participating laboratories reported the same results, indicating the reproducibility and robustness of Y-chromosome SNP typing.
A total of 535 samples from six different European populations were also analysed. In Galicia (NW Spain) and Belgium, the most frequent haplogroup was R1b*(xR1b1,R1b3df). Haplogroup F*(xK) is one of the most frequent in Austria and Denmark, while the lowest frequency appear in Belgium.
Haplogroup frequencies found in this collaborative study were compared with previously published European Y-chromosome haplogroup data.
Link
July 19, 2005
Neighborhood segregation in the United States
Timberlake and Iceland examined all 323 of the nation's metropolitan areas, using Census data from 1970-2000. They searched for trends by exploring four key measures of residential inequality: dissimilarity, entropy, isolation and net difference. The researchers focused on four racial and ethnic groups: Caucasians, African Americans, Asians and Latinos. "African Americans continue to be the most segregated group from whites, but we also found that on average, African Americans have experienced greater declines in segregation," Timberlake says. "So, if that trend continues, Latinos will become the most segregated population by the middle-to-end of the next decade," he says.
Culture switching
In the journal American Psychologist (July 2000), they showed how it might work. Hong Kong students, who are raised in traditional Chinese families but also immersed in Western education and culture, were shown photos of icons that were culturally neutral, American or Chinese before seeking their opinion of what was happening in a photograph of a lone fish swimming ahead of others.
Students who had viewed Chinese icons were more likely to see the lone fish as being chased, while those exposed first to American icons saw the lone fish as a leader.
...
For the new study, Hong and Wong worked with 171 Hong Kong Chinese college students who were faced with the prisoner's dilemma: to cooperate or defect. In such a scenario, each player gains if each cooperates, but only the defector gains more if another player cooperates.
Students were paired with friends or strangers and shown either Chinese or American icons; control groups viewed neutral geometric drawings.
Friends participating after viewing Chinese primes not only were more likely to cooperate, they were much more confident that their partners also would choose cooperation strategies than those shown American icons before facing the problem.
When partners were strangers, those viewing the Chinese primes were only slightly more likely (63 percent to 59 percent) to cooperate with each other, the researchers found.
July 18, 2005
Decline in performance of medical examinees in the UK
After a steady increase between 1985 and 1997, the overall performance of postgraduate students taking the Part 1 examination of the Membership of the Royal Colleges of Physicians of the United Kingdom, MRCP (UK), which is a part of higher specialist training, showed a decline from 1997 until 2001. Performance on specific exam questions that were repeated across the years was 14.1% lower in 2001 than in 1996.The causes of this decline are not obvious:
"The 'dog-leg' in performance [in] 1997 was not an artefact of changed Examination Regulations, mix of UK and overseas candidates, or time from qualification until taking the Examination", write the authors. "Study 2 confirmed that performance in 2001 was significantly worse than in 1996, that the poorer performance was found in graduates of UK medical schools, and that candidates passing the Examination in 2001 performed less well than those passing in 1996."Here are some data on the ethnic composition of medical doctors in the UK in recent decades.
DNA Heritage SNP testing
Also, check out this wonderful demonstration of the distribution of Y-haplogroups around the world. Very convenient as a reference.
July 16, 2005
25,800-year old Australoid from Thailand
The closest sample to the Moh Khiew specimen is the Late Pleistocene Coobool Creek from Australia, and the next closest is the modern Australian Aborigines. The Early to Middle Holocene Southeast Asian samples such as Liang Toge from Flores, Mai Da Dieu from Vietnam and Ban Kao from Thailand are distant from the Moh Khiew specimen.Interestingly, the sample's mtDNA is similar to that of the Semang:
Oota et al. (2001) analysed mtDNA recovered from the Moh Khiew specimen and found continuity with the Semang Negrito foragers living in the Malay Peninsula.
Homo (Article in Press)
A morphometric analysis of the Late Pleistocene Human Skeleton from the Moh Khiew Cave in Thailand
Hirofumi Matsumura and Surin Pookajorn
Abstract
Few Late Pleistocene human remains have been found in Southeast Asia and the morphological features of the people of that age are still largely unknown due to the virtual lack of human remains in the area. Recent excavations at the Moh Khiew Cave in Thailand resulted in the discovery of a Late Pleistocene human skeleton in a relatively good state of preservation. An AMS radiocarbon date on the charcoal sample gathered from the burial gave a result of 25,800±600 BP, implying that the inhabitants of Moh Khiew Cave resided in a part of Sundaland during the last glacial age. In debates on the population history of Southeast Asia, it has been repeatedly advocated that Southeast Asia was occupied by indigenous people akin to present-day Australo-Melanesians prior to an expansion of migrants from Northeast Asia into this area. Morphometric analyses were undertaken to test the validity of this hypothesis. In the present study, cranial and dental measurements recorded from the Moh Khiew remains are compared with those of early and modern samples from Southeast Asia and Australia. These comparisons demonstrate that the Moh Khiew specimen resembles the Late Pleistocene series from Coobool Creek, Australia in both cranial and dental measurements. These results suggest that the Moh Khiew skeleton, as well as other fossil remains from the Tabon, Niah and Gua Gunung sites, represents a member of the Sundaland population during the Late Pleistocene, who may share common ancestry with the present-day Australian Aborigines and Melanesians.
Link
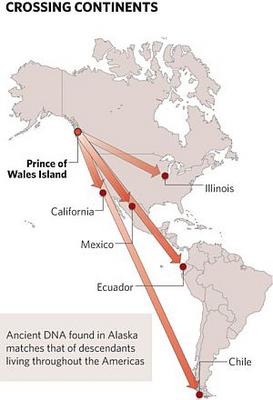
10,300-year old Alaskan had mtDNA haplogroup D
Molecular Clock reset
What is important is that the new calibration leads to significantly lower estimates of events in human evolutionary history, bringing the mtDNA time depth of humanity much closer to the present, and closer to the time depth of the human Y chromosome. Perhaps though, the divergence times of human Y chromosomes may also turn out to be overestimated.
Molecular Biology and Evolution 2005 22(7):1561-1568
Time Dependency of Molecular Rate Estimates and Systematic Overestimation of Recent Divergence Times
Simon Y. W. Ho et al.
Studies of molecular evolutionary rates have yielded a wide range of rate estimates for various genes and taxa. Recent studies based on population-level and pedigree data have produced remarkably high estimates of mutation rate, which strongly contrast with substitution rates inferred in phylogenetic (species-level) studies. Using Bayesian analysis with a relaxed-clock model, we estimated rates for three groups of mitochondrial data: avian protein-coding genes, primate protein-coding genes, and primate d-loop sequences. In all three cases, we found a measurable transition between the high, short-term (<1–2 style="font-weight: bold;">modern humans and Neandertals (354 ka; 222–705 ka), Neandertals (108 ka; 70–156 ka), and modern humans (76 ka; 47–110 ka). If the rate curve for a particular taxonomic group can be accurately estimated, it can be a useful tool for correcting divergence date estimates by taking the rate decay into account. Our results show that it is invalid to extrapolate molecular rates of change across different evolutionary timescales, which has important consequences for studies of populations, domestication, conservation genetics, and human evolution. Link
July 15, 2005
Population structure in Interleukin-13 (IL13) locus
As observed in other studies,19 the East African populations are situated closer to the Eurasian populations in the MDS plot compared to West African populations, and the Chinese population has an intermediate position between Amerindian and European populations (Figure 3).
Genes and Immunity (2005) 6, 53−65
Divergent patterns of linkage disequilibrium and haplotype structure across global populations at the interleukin-13 (IL13) locus
E Tarazona-Santos et al.
Abstract
Interleukin-13 (IL-13) is a cytokine involved in Th2 immune response, which plays a role in susceptibility to infection by extracellular parasites as well as complex diseases of the immune system such as asthma and allergies. To determine the pattern of genetic diversity at the IL13 gene, we sequenced 3950 bp encompassing the IL13 gene and its promoter in 264 chromosomes from individuals originating from East and West Africa, Europe, China and South America. Thirty-one single-nucleotide polymorphisms (SNPs) arranged in 88 haplotypes were indentified, including the nonsynonymous substitution Arg130Gln in exon 4, which differs in frequency across ethnic groups. We show that genetic diversity and linkage disequilibrium (LD) are not evenly distributed across the gene and that sites in the 5' and 3' regions of the gene show strong differentiation among continental groups. We observe a divergent pattern of haplotype variation and LD across geographic regions and we identify a set of htSNPs that will be useful for functional genetic association studies of complex disease. We use several statistical tests to distinguish the effects of natural selection and demographic history on patterns of genetic diversity at the IL13 locus.
Link
The southern origin of O3
American Journal of Human Genetics (Onlin early)
Y-Chromosome Evidence of Southern Origin of the East Asian–Specific Haplogroup O3-M122
Hong Shi et al.
The prehistoric peopling of East Asia by modern humans remains controversial with respect to early population migrations. Here, we present a systematic sampling and genetic screening of an East Asian–specific Y-chromosome haplogroup (O3-M122) in 2,332 individuals from diverse East Asian populations. Our results indicate that the O3-M122 lineage is dominant in East Asian populations, with an average frequency of 44.3%. The microsatellite data show that the O3-M122 haplotypes in southern East Asia are more diverse than those in northern East Asia, suggesting a southern origin of the O3-M122 mutation. It was estimated that the early northward migration of the O3-M122 lineages in East Asia occurred ∼25,000–30,000 years ago, consistent with the fossil records of modern humans in East Asia.
Link
July 14, 2005
Early Modern Humans
The overall impression one gets from this paper is that the human paleoanthropological record is extremely limited. Skulls before 28,000BP are few and are separated by long gaps in space and time. Skulls after 28,000BP are mostly European. Trinkaus criticizes molecular biology for its wider confidence intervals and unstated theoretical assumptions, but one wonders whether the situation in the traditional osteological approach is much better, and whether pronouncements on what constitutes archaic vs. modern can be so easily done with such meagree material. For example, the Herto skulls were assigned to a new human subspecies Homo sapiens idaltu, but it was later found that the more "modern" Omo remains were about 50,000 years older. So, there is no clear progression between archaicity to modernity, even in eastern Africa - not to mention that there is a huge gap between these early remains and the multiple (mostly European) skulls dating to post-28,000BP times.
Trinkaus pronounces both the replacement and regional continuity theories dead. Humans are not descended from regional populations of Homo, and they are not descended from a single African population. Rather, he prefers the assimilation model, whereby a population from the east African ecological zone expanded, assimilating to some extent archaic human populations.
Annual Review of Anthropology
First posted online on June 14, 2005
Early Modern Humans
Erik Trinkaus
Perceptions of the emergence and spread of modern humans have changed recently through the reanalysis of fossils, an improved geochronological framework, and the discovery of a few specimens. Early modern humans in various portions of the Old World exhibit complex and varying mosaics of archaic, modern, and regional morphological characteristics. On the basis of this pattern, in conjunction with the emerging chronology of the earliest modern humans, the paleontological data indicate an assimilation model for modern human origins, in which the earliest modern humans emerged in eastern Africa, dispersed briefly into southwestern Asia, and then subsequently spread into the remainder of Africa and southern Asia, eventually into higher latitude Eurasia. The earliest modern humans outside of the core area of eastern Africa can be understood only if a variable degree of admixture with regional groups of late archaic humans occurred. Current and expected fossil and molecular data are unlikely to illuminate the degree of assimilation that took place in most regions of the Old World. However, the current chronological and phylogenetic framework provides the basis for ongoing investigation of the nature of this Late Pleistocene transitional period.
Link
Ethnoancestry
Let's take haplogroup J, for example. Someone who belongs to it can learn that he is either J1 or J2 with an extremely small probability that he will turn out to be J*(xJ1, J2). On the other hand, some haplogroups such as R1a don't have a strong substructure (yet) as defined by known SNPs, so it is not entirely clear whether one should wait for more research to be published on the matter, i.e., to figure out which SNPs are interesting, widely prevalent, and worth testing for.
July 13, 2005
Lactose malabsorption, climate, and cattle disease
Evolution and Human Behavior
Volume 26, Issue 4 , July 2005, Pages 301-312
Dairying barriers affect the distribution of lactose malabsorption
Gabrielle Bloom and Paul W. Sherman
Abstract
Most mammals stop drinking milk at weaning, which is also the time when they cease producing lactase, the digestive enzyme that hydrolyzes lactose. Cessation of lactase production and milk drinking also characterize most human populations, especially those of African and Asian descent. However, a genetic mutation that maintains the functionality of lactase production into adulthood occurs commonly among populations from northern Europe, where dairying is practiced routinely. Indeed, the ability to absorb lactose is nutritionally beneficial for adults only if milk consistently is available. What determines the distribution of dairying? We hypothesized that specific environmental circumstances affect where milk-producing ungulates can be raised safely and economically, thus influencing the geographical occurrence of dairying and lactase persistence. To evaluate this hypothesis, we compiled data on adult lactose absorption (LA) and malabsorption (LM) frequencies in 270 indigenous African and Eurasian populations (Appendix A). Partial correlation analyses revealed that, as predicted, adult LM is associated with extreme climates (at high and low latitudes) and, more significantly, with the historical (pre-1900) geographical occurrence of nine deadly, communicable diseases of cattle. These results suggest that areas where adult LM predominates are those where it is impossible or dangerous to maintain dairy herds.
Link
July 07, 2005
Archaic mtDNA in Taiwan and Polynesian origins
PLoS Biology (Online)
Mitochondrial DNA Provides a Link between Polynesians and Indigenous Taiwanese
Jean A. Trejaut et al.
Genetic affinities between aboriginal Taiwanese and populations from Oceania and Southeast Asia have previously been explored through analyses of mitochondrial DNA (mtDNA), Y chromosomal DNA, and human leukocyte antigen loci. Recent genetic studies have supported the “slow boat” and “entangled bank” models according to which the Polynesian migration can be seen as an expansion from Melanesia without any major direct genetic thread leading back to its initiation from Taiwan. We assessed mtDNA variation in 640 individuals from nine tribes of the central mountain ranges and east coast regions of Taiwan. In contrast to the Han populations, the tribes showed a low frequency of haplogroups D4 and G, and an absence of haplogroups A, C, Z, M9, and M10. Also, more than 85% of the maternal lineages were nested within haplogroups B4, B5a, F1a, F3b, E, and M7. Although indicating a common origin of the populations of insular Southeast Asia and Oceania, most mtDNA lineages in Taiwanese aboriginal populations are grouped separately from those found in China and the Taiwan general (Han) population, suggesting a prevalence in the Taiwanese aboriginal gene pool of its initial late Pleistocene settlers. Interestingly, from complete mtDNA sequencing information, most B4a lineages were associated with three coding region substitutions, defining a new subclade, B4a1a, that endorses the origin of Polynesian migration from Taiwan. Coalescence times of B4a1a were 13.2 ± 3.8 thousand years (or 9.3 ± 2.5 thousand years in Papuans and Polynesians). Considering the lack of a common specific Y chromosomal element shared by the Taiwanese aboriginals and Polynesians, the mtDNA evidence provided here is also consistent with the suggestion that the proto-Oceanic societies would have been mainly matrilocal.
Link
July 05, 2005
40,000-year-old footrpints from the Americas
However, both lines of evidence do not exclude the possibility of earlier habitation, provided that it was limited enough in scope. Perhaps early humans did arrive to the New World but did not manage to successfully colonize it and create a lasting presence. If that is the case, then we would not expect to find traces of the genes of these early colonists among living Native Americans, and the archaeological evidence would be meagre, or absent.
The new discovery does not of course prove that man arrived to the New World at an early age, but hopefully it may trigger more research into the earlier geological time periods, which were hitherto considered to be archaeologically barren.
July 04, 2005
Journal of Genetic Genealogy
July 03, 2005
The declining rate of technological innovation
Dr. Huebner proposes that technological innovation is limited both physically (e.g., it is impossible to transcend the limitations of natural laws), but also economically (it may be possible, but not practical to do something). According to his interpretation, the observed trend due to economic limitations, although he does leave open the possibility that it may be cognitive, i.e., that the human mind may have been pushed close to its limit.
Technological Forecasting and Social Change (Article in Press)
A possible declining trend for worldwide innovation
Jonathan Huebner
Abstract
A comparison is made between a model of technology in which the level of technology advances exponentially without limit and a model with an economic limit. The model with an economic limit best fits data obtained from lists of events in the history of science and technology as well as the patent history in the United States. The rate of innovation peaked in the year 1873 and is now rapidly declining. We are at an estimated 85% of the economic limit of technology, and it is projected that we will reach 90% in 2018 and 95% in 2038.
Link
July 02, 2005
Scanning the genome for intelligence genes
American Journal of Human Genetics (Online Early)
A Genomewide Scan for Intelligence Identifies Quantitative Trait Loci on 2q and 6p
Danielle Posthuma et al.
Between 40% and 80% of the variation in human intelligence (IQ) is attributable to genetic factors. Except for many rare mutations resulting in severe cognitive dysfunction, attempts to identify these factors have not been successful. We report a genomewide linkage scan involving 634 sibling pairs designed to identify chromosomal regions that explain variation in IQ. Model-free multipoint linkage analysis revealed evidence of a significant quantitative-trait locus for performance IQ at 2q24.1-31.1 (LOD score 4.42), which overlaps the 2q21-33 region that has repeatedly shown linkage to autism. A second region revealed suggestive linkage for both full-scale and verbal IQs on 6p25.3-22.3 (LOD score 3.20 for full-scale IQ and 2.33 for verbal IQ), overlapping marginally with the 6p22.3-21.31 region implicated in reading disability and dyslexia.
Link
