Figure S3 contains the results of the STRUCTURE analysis (from K=2 top row). The broad results are consistent with similar past analyses, but many populations that were previously not examined in the global context are included. You can probably discern many interesting features with your magnifying glass, but I will limit myself to a handful:

- Greeks from northern Greece (#15) and Cyprus (#9) appear fairly unremarkable in their genetic makeup; indeed most Europeans appear similar in this broad global context
- The small sample of 4 Turks (#38) shows a small membership in "Asian" clusters, although these appear to be mostly of the "Central/South Asian", rather than the "East Asian" variety. This probably makes them similar somewhat to the Adygei from the Caucasus in a previous analysis. This element does not, however, seem to be very important in Near Eastern Semitic populations included in the HGDP panel, so it would be interesting to see how the transition from European to Central/South Asian Caucasoids occurs in Transcaucasia, Iran, and the various -stans.
- CEU Utahns seem to lack the "purple" component altogether (bottom row), and in this they are most similar to Britons and Iberians, perhaps signifying a peculiarity of Western-most populations in Europe.
Table S5 shows the percentage of haplotypes shared between different European regions and the African Yoruba (YRI) sample.
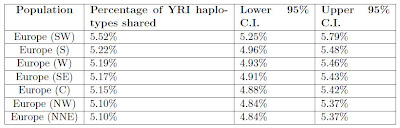 Most European regions are within 0.12% of each other, but Southwest Europe has an elevated percentage of 5.52% of 0.3% higher than the next highest percentage. Thus, the hypothesis of a separate influence in Iberia from Africa that did not pass from east-to-west seems reasonable.
Most European regions are within 0.12% of each other, but Southwest Europe has an elevated percentage of 5.52% of 0.3% higher than the next highest percentage. Thus, the hypothesis of a separate influence in Iberia from Africa that did not pass from east-to-west seems reasonable.However, the STRUCTURE analysis (see above) does not indicate a substantial presence of the Yoruba (dark green) element anywhere in Europe, consistent with the recent paper of Tishkoff et al. It appears more likely that the similarity is due to a (as of yet unsampled) element common to both populations, perhaps of North African or other intermediate origin.
In conclusion, this is a very interesting paper (of great interest for South Asia and the Americas as well, not covered in my post), as it furthers our understanding of the global distribution of genetic diversity. It would be wonderful to combine such a comprehensive global dataset with the substantial African one of the Tishkoff paper, but unfortunately the different types of markers genotyped make it at present impossible.

15 comments:
Why are some of the European countries showing more of the East Eurasian blue on the "Mexican admixture" plot and much less or none at all on the Global admixture plot? For example, Italy...
@Polak:
As far as I can see, the thin (purple?) upper component seen in Europeans at K=7, as well as the green one most visible at K=6 are minor components with no regional classification at all.
The regional components are all visible at K=5: Red-West Eurasian, Blue-East Asian, Black-Yoruba, Green-South Asian and Yellow-Native American. At that level basically Turkey (and to a lesser extent Cyprus and some SE European samples, such as Italy) show a minor but noticeable South Asian component among West Eurasians.
The "Mexican Admixture" graph intead only reaches to K=4 and the sample is different with no South Asians. Only Turkey shows a very small, yet visible, East Asian factor in their genetic pool.
I think you're right about K5 being the one that counts in terms of admixture. That would make sense and fit in with the "Mexican admix" plot.
By the way, would you say the purple is some kind of ancient Eurasian component that has changed in frequency here and there via drift?
Or is that just the European markers starting to break up after K5...I think so, eh?
Don't know what to reply, really. The only thing pretty much obvious is that it is alternative to the orange component (first seen at K=6), so basically must mean the same.
But what confuses me most is that this one and the green component seem to increase (and not decrease, as would be logical) as we go deeper in the K means structure. I mean: the green cmponent appears notably more common among Europeans at K=6 than at K=4 or K=5, while the orange component appears more common among East Asians (and only among them) at K=7 than at K=6.
I really can't make much sense of that. Whatever the case, at K=7, the purple/orange component(s) are similarly present among all Eurasians, though, as Dienekes points out some have more purple and others more orange.
Unlike him, I seem to see as more orange/less purple the following samples: CEU, Croatia, Bosnia, Portugal and Swiss-Italians. And in Asia: CHB, JPT and most Indians. The most markedly purple samples instead are: Germany, Greece, Russia, Swiss-German and Swiss in general. Do you make any sense of it? Would it be Europe-only, I might think of some IE or maybe Jewish (Khazar?) component (the purple one) but as they are shared with remote locations as Japan and India where such explanations cannot stand... meh! Who knows?
Maybe just something pan-Eurasian, very old stuff as you say.
Maju,
Well after the Hun (and allies) army was defeated in the Champagne district of France, by the Roman (and allies) army, some Huns settled in Champagne, but many settled in some of the mountainous Swiss valleys.
Here's an article:
http://www.simaqianstudio.com/forum/index.php?showtopic=5044&mode=threaded&pid=60330
Which states that the people of the Anniviers valley (aka Val d'Anniviers or Eifischtale) were of Hunnish descent, and spoke a dialect of Hungarian, with similarities to the Szeklers, of Hungary.
So that might be the purple component?
Also the Avars settled in the Balkans, and the island of Hvar, off the Croatian (or Dalmatian) coast, is high in Central Asian DNA, so some may have found it's way into Greece from there?
Only 4 individuals hardly represent a complex and diverse population like Turks. I could just ignore the results for Turks without making further comments on them.
Even if I assume for a minute that they are a good representative of the whole Turkish population, it is still too early to make judgements about the genetic structure of Turks in comparison with the rest of the world, because, as Dienekes noted, we should also see large samples of the populations of Transcaucasia, Iran and the former Soviet republics of Central Asia in STRUCTURE analyses.
As far as my eyes can see, the East Asian component of Turks is negligible compared to the South Asian one in the global plot. So the clearly visible East Asian component of Turks in the Mexican admixture plot may actually be mostly South Asian rather than East Asian.
Ok with Onur. I said it vas "a very small, yet visible, East Asian factor". Agreed that the SA component is strongest if anything, not just among Turks - what seems absolutely logical to me considering the many haplogroups shared east and west of Balochistan and my impression that the regional "borders" were "closed" earlier for East Asia and Sahul than between West and South Eurasia.
As for PConroy:
Maju,
Well after the Hun (and allies) army was defeated in the Champagne district of France, by the Roman (and allies) army, some Huns settled in Champagne, but many settled in some of the mountainous Swiss valleys.
There are other such cases: in Croatia and Italy and surely even more in Germany and the like (not registered anywhere) but that doesn't seem to mean anything, as we should see it related to either the East Asian component or something that should be meaningful in Turkey (Huns were Turkic after all).
My impression is that the minor purple component has highest density in West Eurasia in fact, if anywhere. Whatever that means.
Also for some reason it does not appear in any of the 4 standard samples (CEU, CHB, JPT and YRI), while it does appear in similar samples of the study, what I suspect that must mean that such samples are all affected by some systematic distortion somehow.
Okay, Maju was right...
There's no point going past K5 when looking at inter-continental admix there, because after that the local clusters start breaking up, noise starts creeping in, and it's hard to gain anything of value at that point.
Also, the reason the CEU, CHB and JPT don't have any purple is because they were "done" differently to the rest, which come from POPRES.
Mystery solved. :p
There's no point going past K5 when looking at inter-continental admix there, because after that the local clusters start breaking up, noise starts creeping in, and it's hard to gain anything of value at that point.Then that answers your initial question why some European countries show more blue component on the Mexican admixture plot than on the global plot: The blue component of Europeans on the Mexican plot is almost totally South Asian (as they aren't represented), rather than East Asian. --> See the transition from K=3 to K=4 on the global plot.
The same is true for Turks, as there is no East Asian component in them, but a relatively strong South Asian one on the global plot.
But, as I emphasized before, only 4 individuals CANNOT represent Turkey (or any other country for that matter).
I'm not questioning that in principle, although, as they took local wifes and concubines (and co-opted local men maybe), their East Asian blood should have got diluted to near-nothingness in rather few generations. And more after they settled among other locals and lost their distinctive identity.
Additionally, in the wide picture Turco-Mongol peoples (Huns included but many ore after them) were much more influential in Eastern Europe, from Hungary and Rumania towards the east. Still their genetic impact in West Eurasia is overall very limited, even among Turkic-speaking peoples.
The minor purple component might (only might) be some sort of deeply old Eurasian component, with some tendency towards the north of the continent (in contrast with the minor orange one that may be more southwardly distributed). But it is not specifically East Asian, much less when the populations with greatest density of that component are in Europe.
The design is really good one and eye catching...Thanks for sharing....
Generic Viagra, Kamagra Online, Generic Levitra
Post a Comment