An interesting table from the paper shows the number of significant principal components within each continent:
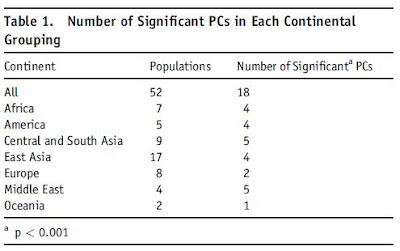
Notice the large number of principal components in the Middle East and Central Asia, suggesting a very significant differentiation in these regions, despite the small number of tested populations, and underscoring the need for more comprehensive sampling.
In the case of the Middle East, where only Afroasiatic populations were sampled, this is even more remarkable; further study of Indo-European, Turkic, and Caucasian speakers from the region will no doubt reveal further differences among them. As for Central Asia, the large number of significant principal components is related to both the inter-racial difference between Caucasoids and Mongoloids in this contact zone, as well as intra-racial differences.
The supplementary material (pdfs) has plots for the significant components of the seven continental regions. Progress will now occur by extending population sampling to unexamined populations and by full-genome sequencing which will allow us to detect finer-level distinctions even in fairly homogeneous regions of the world such as Europe.
American Journal of Human Genetics doi:10.1016/j.ajhg.2009.04.015
Genome-wide Insights into the Patterns and Determinants of Fine-Scale Population Structure in Humans
Shameek Biswas et al.
Abstract
Studying genomic patterns of human population structure provides important insights into human evolutionary history and the relationship among populations, and it has significant practical implications for disease-gene mapping. Here we describe a principal component (PC)-based approach to studying intracontinental population structure in humans, identify the underlying markers mediating the observed patterns of fine-scale population structure, and infer the predominating evolutionary forces shaping local population structure. We applied this methodology to a data set of 650K SNPs genotyped in 944 unrelated individuals from 52 populations and demonstrate that, although typical PC analyses focus on the top axes of variation, substantial information about population structure is contained in lower-ranked PCs. We identified 18 significant PCs, some of which distinguish individual populations. In addition to visually representing sample clusters in PC biplots, we estimated the set of all SNPs significantly correlated with each of the most informative axes of variation. These polymorphisms, unlike ancestry-informative markers (AIMs), constitute a much larger set of loci that drive genomic signatures of population structure. The genome-wide distribution of these significantly correlated markers can largely be accounted for by the stochastic effects of genetic drift, although significant clustering does occur in genomic regions that have been previously implicated as targets of recent adaptive evolution.
Link

21 comments:
Notice the large number of principal components in the Middle East and Central Asia, suggesting a very significant differentiation in these regions...
But please compare with the supplemental data table S2: the permutation method gives highest number to America and the ANOVA method to East Asia.
... despite the small number of tested populations, and underscoring the need for more comprehensive sampling.
Absolutely. No Australian Aborigines, no Indians and no East Africans. You cannot draw any global conclusions from that.
"Absolutely. No Australian Aborigines, no Indians and no East Africans. You cannot draw any global conclusions from that".
Largely true. But you can draw a few conclusions. I was interested to see the distinction between Papuans and Melanesians. Most of us are inclined to think of them as a single group even though mtDNA and Y-chromosome suggests otherwise.
"Notice the large number of principal components in the Middle East and Central Asia".
Perhaps because humans have actually been there for a very long time? Maybe they moved through the Iranian Plateau to India rather that along the coast.
terryt said,
"I was interested to see the distinction between Papuans and Melanesians. Most of us are inclined to think of them as a single group even though mtDNA and Y-chromosome suggests otherwise."
I believe previous studies of autosomal genetic and phenotypic variation have found highland Papuans to be more similar to Australian aborigines than they are similar to coastal Papuans or Melanesians. I previously have posited a hypothesis that highland Papuans might have better preserved the genes of the earliest settlers of Sahul, whereas coastal Papuans and Melanesians might be derived from a different isolated Southeast Asian stock that has expanded into Near Oceania during the dispersal of Austronesian languages.
Thanks Ebizur. That supports the idea that there were several movements across Wallace's line rather than the current diversity being a product of a single expansion containing all surviving groups.
"coastal Papuans and Melanesians might be derived from a different isolated Southeast Asian stock that has expanded into Near Oceania during the dispersal of Austronesian languages".
I realised after I'd responded what you'd actually said. For several reasons it would be difficult to argue that the difference between highland and coastal Papuans is solely because of the Austronesian expansion. The haplogroups involved in the Austronesian expansion seem to be different to those characteristic of most coastal Papuans.
To start with the Polynesian language is a subset of Austronesian, and Polynesians are almost exclusively mtDNA B and Y-haps C2 and O3. So at least the Polynesian branch of Austronesian is associated with these three haplogroups. The same haplogroups, along with several Austronesian languages, are found scattered through Melanesia, and so the Austronesian languages and the above haplogroups are probably associated there too. It's even possible that the same haplogroups are in fact associated with the Austronesian-related languages in SE Asia.
Another difficulty in associating the differences between New Guinea and Melanesia as being simply a product of the Austronesian expansion is that many languages in Melanesia are not actually Austronesian. They are lumped into an Indo-Pacific group, but that doesn't necessarily mean they are at all related to each other. But they are not Austronesian-related. So, the Austronesian expansion is not responsible for the difference between coastal and highland Papuans. Some other separate migration from SE Asia is required to account for the difference.
When we go further back we come to another very interesting fact. The vast majority of Australian mtDNA haplogroups are N-derived but if we take out the (probably) Austronesian-related B, and the P common to both Australia and New Guinea, we find New Guinea has a majority of M-derived haplogroups. And when we look at Y-chromosomes we find that more than 60% of Australian haplogroups are C-derived and the vast majority of Y-haps in the New Guinea highlands are F-derived. This suggests that these two regions were not originally settled by the same population. So that leaves us with at least four Wallacea crossings from west to east.
terryt,
I think there has been some misunderstanding. My hypothesis is that the ancestors of the Melanesians have arrived to their current locations (i.e. the islands of Melanesia and some coastal parts of New Guinea) as part of the Austronesian expansion, not that they are necessarily the "purest" descendants of the original speakers of Proto-Austronesian.
I can't quite understand what you mean by that Ebizur. The Northern Solomons and New Britain/New Ireland (all part of Melanesia) were already occupied by the time of the Austronesian expansion, although I'd agree that Austronesian-speaking people were probably the first people into the Southern Solomons, Vanuatu and New Caledonia. The Austronesians had largely bypassed the northern people although the people from further north then followed along, eventually reaching as far as Fiji.
terryt,
"I can't quite understand what you mean by that Ebizur. The Northern Solomons and New Britain/New Ireland (all part of Melanesia) were already occupied by the time of the Austronesian expansion..."
It doesn't really matter whether the islands were "already occupied" or not. That reminds me too much of the old "R1b must be descended from Palaeolithic Cro-Magnons of Western Europe because it is most common in present-day populations of the Iberian Peninsula and the British Isles" fallacy.
Besides the fact that most Melanesians and many coastal New Guineans speak Austronesian languages, there is also the crucially important (yet often overlooked of late, perhaps for the sake of political correctness) issue of phenotypic variation. Melanesians tend to look different from Papuan highlanders or Australian aborigines, who are probably the best candidates for the indigenous populations of Sahul.
Furthermore, if I am not mistaken, some examples of Y-DNA haplogroup M have been found in places as far away as Malaysia. Can you tell me the estimated age of Karafet's newly restructured Y-DNA haplogroup M? I don't recall what the estimated age for this one was (if I have ever seen it), but I suspect that it might not be too old.
Signatures of positive selection in genes associated with human skin pigmentation as revealed from analyses of single nucleotide polymorphisms
"Our results showed that African populations tended to carry the ancestral alleles of the studied SNPs...and that human populations with dark skin colour tended to cluster together in the MDS and STRUCTURE analyses. The clustering of the Bougainville Islander population reflects the fact that this relationship is not (or at least not solely) caused by geography but also by the underlying pigmentation genes. Bougainville Islanders from Papua New Guinea are known to be one of the most highly pigmented people in the world...In our results they appeared to be closer to Sub-Saharan Africans populations - with whom they share the dark skin colour phenotype - than to the second Papuan New Guinea sample in the dataset, with whom they share their recent population history, as observed in several datasets based on neutral genetic variation from autosomal, Y-chromosomal and mtDNA analyses...."
Why are Bougainville Islanders closer to Sub-Saharan Africans populations than their fellow Melanesians for "Signatures of positive selection in genes associated with human skin pigmentation"
Have they dark skin to protect from strong UV that the nearest Melanesians (in other Solomon Islands) are not exposed to? Are they geographically isolated NoBougainville Island has an abundance of land suitable for women to support themselves by w garden agricuture. Hence polygyny became viable
Origins of black Africans "Relaxed sexual selection of women may also explain why, among sub-Saharan Africans, skin color is visibly darker in high-polygyny agriculturalists than in low-polygyny hunter-gatherers (i.e., Khoisans, pygmies) even though both are equally indigenous (Bourguignon and Greenbaum, 1973, pp. 171-175; Cavalli-Sforza, 1986a; Cavalli-Sforza, 1986b; Manning et al., 2004; Weiner et al, 1964). Skin color does, in fact, influence mate choice in all human societies; generally speaking, men prefer women who are lighter-skinned than the population mean (van den Berghe and Frost, 1986). In sub-Saharan societies, the preference is for so-called 'red' or 'yellow' women (see earlier post). Wherever African men were less able to act on this preference, there would have been less selection for lighter-skinned women and thus less counterbalancing of selection for darker skin to protect against sunburn and skin cancer (Aoki, 2002; Frost, 2007; Frost, 1994).
This preference may also have been crowded out by other mate-choice criteria. Vilakazi (1962, pp. 59-60) states: "The traditional Zulu does not make physical beauty a first priority or even an important qualification in a wife; and the skin colour of the woman is of little importance." In a rating study, Dixson et al. (2006) examined mate-choice criteria among subsistence farmers in Bakossiland, Cameroon, including preferred skin color of a potential female partner. No consistent preference emerged. This ambivalence was noted by Ardener (1954, p. 72) among the Ibo of Nigeria:
In the choice of a wife, yellow-skinned girls are regarded as beauties, and, other things being equal, they command higher bride prices. On the other hand it is generally held, especially by dark-complexioned persons, that yellow-skinned people are not as strong as the dark and do not live as long. A 'black' girl is said to be a harder worker. … A Mission headmaster was of the opinion that the preference for yellow girls was greater nowadays than in his youth. He thought that the reason for this was that people formerly looked for strength rather than beauty and tended to marry black girls. He claimed that black people had greater powers of endurance, and he cited his own village where, he said, of the oldest six or seven people, only one was yellow.
In Kenya, McVicar (1969, p. 242) notes similar views on the merits of ‘black’ versus ‘brown’ wives: "Among these tribes black girls are usually regarded as hard workers, possibly because many consider themselves fortunate enough to be married." In traditional African societies, women had to produce enough food for the entire family, typically through hoe farming in the sun. There was thus a premium on darker women. Lighter women may have been preferred aesthetically, but this preference remained unexpressed."
@Ebizur:
Restructured M should be older than M1 (old M) by mere logic. For me M may well be one of the first K-derived expanding lineages, what would make it near 50-60 kya.
Karafet did not even bother estimating the age of M but her age estimates are based on the assumtion that CF (i.e. C'D'E'F) being as young as 70 kya, which is an arbitrary assumption.
You argue (elsewhere but is implicit here too) that C2, being present in parts of Indonesia would be Austronesian by origin. I sincerely doubt it, as Ausronesians are original from Taiwan and then Luzon, not Indonesia nor Malaysia. C2 and other "Melanesian" clades would have just joined them as they migrated, logically.
I am with Terry in this case.
http://epress.anu.edu.au/austronesians/austronesians/mobile_devices/ch09s05.html
"...The Polynesian groups have a position intermediate between northern China and coastal Melanesia. In the phylogenetic analysis, Melanesians from the north Papua New Guinea coast (Madang) cannot be discriminated from the Tolai of New Britain and these groups have a DR, DQ profile similar to that in Melanesians from New Caledonia. The Fijian sample reflects some admixture with Polynesian elements and is equidistant to New Caledonia and Western Samoa in the eigenvector diagram. Micronesians from Nauru and Kiribati are well-separated from Polynesians due to a high frequency of DRB1*1202, a DR5 allele that is commonly found in Javanese, less frequently in southern Chinese, and rarely elsewhere. Non-Austronesian-speaking Melanesians from the New Guinea Highlands show closer affinity with Australian Aborigines than with any of the Austronesian-speaking Melanesian groups. The Australian Aboriginal populations cluster together, even though about 50 per cent of HLA-DR alleles are unique to Kimberleys Aborigines."
This is one example from a study of immunological genetics, but there is plenty of evidence that Austronesian-speaking Melanesians are more closely related among themselves than they are related to non-Austronesian-speaking New Guinea highlanders or Australian aborigines.
A somewhat related question: what are the Y-DNA and mtDNA haplogroups found among the Aetas in the Philippines?
"Spreading concomitant with the dispersal of Austronesian languages" does not even suggest anything about "the purest descendants of the original Proto-Austronesian speakers." How many times must I repeat this?
But isn't that because of the Lapita phase, that preceded the main Polynesian expansion. Whatever the case Polynesians are not the only Austronesians and you can't construct a theory for the whole based only in the part. It'd be like arguing on the origin of Indoeuropeans based only in Bengalis or Jamaicans, for example.
I haven't said anything about arguing the origin of the Proto-Austronesians. In fact, I have repeatedly asked you and terry to leave off that subject, as it is irrelevant to my proposition.
"Melanesians tend to look different from Papuan highlanders or Australian aborigines, who are probably the best candidates for the indigenous populations of Sahul".
I realised later that I'd answered my own question. Unfortunately I'd considered just New Britain/New Ireland as Melanesia whereas the study obviously includes Melanesians from further south, who are definitely more admixed with Austronesians.
"The Polynesian groups have a position intermediate between northern China and coastal Melanesia".
And Polynesians are presumably basically a hybrid between the people from north (or southern anyway) China and the people from Melanesia. The remainder of Ebizur's quote shows that, as a result of all the to and fro movements across the Pacific, we've finished up with a cline from marginal Polynesia in the east to Melanesians in the west, Chinese and Japanese in the north and Australian Aborigines in the south. We can even extend the cline further west into India. But even in the geographic extremities the people have by now become fairly mixed. In fact the original study appears to indicate such clines are the norm for human differences across the earth.
"Can you tell me the estimated age of Karafet's newly restructured Y-DNA haplogroup M?"
Soory. No.
"I suspect that it might not be too old".
Same here. It's mainly confined to northern Melanesia and as far as I know that region was first settled around 30,000 years ago. So I would guess it would be nearer that age than 'near 50-60 kya'.
"some examples of Y-DNA haplogroup M have been found in places as far away as Malaysia".
My guess (just that) would be that Malaysian Y-hap M would be the product of a movement back west across Wallacea rather than being part of a remnant population.
"what are the Y-DNA and mtDNA haplogroups found among the Aetas in the Philippines?"
A good question. If you find the answer please post it.
This may be useful though fairly old. I haven't time to look through it now:
http://www.pubmedcentral.nih.gov/articlerender.fcgi?artid=1715355
Sorry. That last link is noit very informative now. Thia is intersting though:
http://blogs.nationalgeographic.com/blogs/genographic/2008/12/meeting-the-ati-peoples-in-the.html
Seems Spencer Wells in December last year set out to examine Aeta haplogroups. So we can expect the results any day. A couple of comments in the article seem strange though.
"The first of these, the initial settlement of Southeast Asia more than 40,000 years ago, involved hunter-gatherers walking across now submerged land bridges connecting the islands of the Sunda Shelf".
I understand that the Philippines have never been actually connected to either Sahul or Sunda, so technically they are part of Wallacea. Humans must have reached them by boat.
"the Aeta look more African".
Well. Not the photos I've seen of them, and not the one Wells provides. Perhaps like Timorese.
"The earlier trip with Dr. Rand Allingham yielded data from more than 100 Aeta, and the genetic results we have obtained are fascinating. We're currently writing them up, and don't want to discuss them in too much detail until they have been published, but the Aeta certainly seem to be a mix of both Austronesian and earlier genetic lineages, and their Y-chromosomes appear to be almost completely pre-Austronesian".
So we can look forward to some interesting revelations, unless what Wells has discovered doesn't fit his own theories.
Fsts for Finns :
Finns (Helsinki) :
Austria 0.0060
Bulgaria 0.0090
Czech Republic 0.0060
Estonia 0.0040
France 0.0080
Northern Germany 0.0060
Southern Germany 0.0060
Hungary 0.0060
Northern Italy 0.0130
Southern Italy 0.0160
Latvia 0.0070
Lithuania 0.0070
Poland 0.0060
Russia 0.0060
Spain 0.0110
Sweden 0.0050
Switzerland 0.0090
CEU 0.0060
Finns (Kuusamo) :
Austria 0.0130
Bulgaria 0.0150
Czech Republic 0.0120
Estonia 0.0090
France 0.0150
Northern Germany 0.0120
Southern Germany 0.0130
Hungary 0.0130
Northern Italy 0.0200
Southern Italy 0.0230
Latvia 0.0130
Lithuania 0.0130
Poland 0.0120
Russia 0.0120
Spain 0.0170
Sweden 0.0110
Switzerland 0.0150
CEU 0.0130
As we can see there is less genetic distance, for example, between Northern Italians and Palestinians (fst 0.0108) than between Northern Italians and Finns ( 0.0130-0.0200) or even Latvia (0.0120) or Lithuania (0.0110) ! Bad news for these stupid euro-nationalists that would like to match "political" europe borders with genetic ones ...
Haha! As we can see: Finnish from Helsinki are closer to any other Europeans than Finnish from Kuusamo are to anybody else but Estonians (though Helsinkians are as close to Bulgarians as that).
Bad news for nutheads with an agenda everywhere, I guess. Not even the ethno-political borders of Finland can save your biased agenda from factual data. Not that never ever any nation's borders (a military imposition) had anything to do with geneteics but anyhow...
Everyone knows Finns are outliers in Europe genetically, so are Greeks in general. No big deal there, just signs of the closeness of Greeks to the Middle East, and Finns to Ob-Ugrians and other "Asians". Sardinians tend to cluster apart also. It is a large island after all and mountainous. Lots of places to have remnant populations, and odd mixtures of various European strains. North Italians are basically like run of the mill Central Europeans, nothing much special about them.
I was trying to understand what Ken was waffling about. Melanesians is just a name for black skinned South Pacific Islanders. Doesn't suggest close genetic ties. Papuans are not really Melanesian, not like most Fijian Islanders or the inhabitants of Tanna. Bougainvilleans refer to Papuans their fellow citizens of Papua/New Guineas as Redskins, a derogatory term, as they see Papuans are red coloured, and hence weak. New Guinea and Bougainville have separate genetic histories until linked by Europeans during the Colonial era. They don't have to have similar skin pigmentation. New Guinea is mountainous, high rainfall, and tropical. Mostly it is covered in mists. Having a black skin is not that important in that climate and topography.
Post a Comment