- fastIBD was run with default parameters over a dataset of 512 individuals/264,539 SNPs
- fastIBD identifies segments of relatively recent origin that are shared by individuals. These results should not be construed as measures of overall genetic similarity or origins. Rather, they suggest which populations have exchanged genes in the relative recent past, say, the last two thousand years or so.
- I included all Ashkenazi_D and North_African_Jews_D samples; of the other Dodecad and reference populations, I took random samples of 10 each; running time of fastIBD increases with the square of the number of individuals, so doing this allowed me to run this in less than a day as opposed to about a week.
- Spreadsheet of numeric results, showing sharing (in centi-Morgans, cM)
- Population-level graphical results, showing an ordering of other populations based on mean IBD sharing.
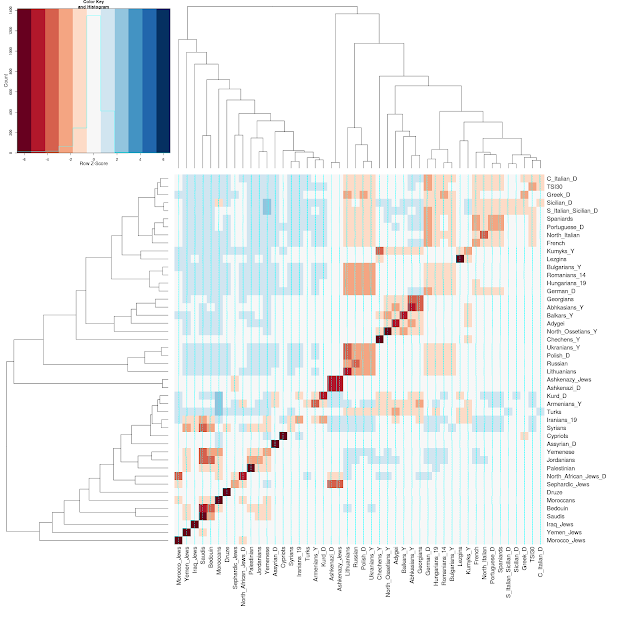
And, here are a couple of the visualizations for a few Jewish populations:
Note that all sources of data are listed on the bottom left of the Dodecad blog.






4 comments:
Really interesting. There is a lot to mull over here.
How is that Sephardic Jews share a larger IBD with Ashkenazic Jews than with each other?
Sephardic Jews are very diverse compared to Ashkenazi Jews. A similar finding was reported in the recent Graham and Coop paper in which Germans shared more IBD with Poles on average than with each other.
The best way to think about this is by drawing a trapezoid like this:
http://en.wikipedia.org/wiki/File:Trapezoid.svg
if AB is much longer than CD, and h is small, then random points in AB are usually further apart than they are to CD.
So, populations with reduced genetic diversity (such as Balto-Slavs, a very recent population expansion, or AJ, a quite recent population isolate) tend to IBD share a lot.
Interesting to see that Morocco Jews How share a larger IBD with non-Jewish Moroccans than with some Ashkenazi, for example. It confirms what was found by the study:
"Admixture occurred
with North African non-Jewish populations. However, as
demonstrated by the apparent unrelatedness of the Jewish and
non-Jewish North Africans, these events were not recent."
Post a Comment