(Last Update Jun 10)
More on this paper as soon as I read it carefully.
Nature News has an overview of the research. This study addresses my
question about the extent of Southern vs. Central/European ancestry in Jews.
It is also entirely consistent with my theory that Diaspora Jews are to a certain extent descended from Italian-Balkan-Anatolian groups, among which they lived in Hellenistic-Roman-Late Antique times; my guess is that Middle-Eastern Jews form a distinct group in relation to European/Syrian ones because, unlike them, they had a smaller opportunity of absorbing Euranatolians, and their admixture -if any- came from linguistically (and probably genetically) related Semitic groups.
UPDATE I (Jun 4)
It is a bit frustrating how the authors did not limit themselves to HGDP but also included a wide variety of populations from POPRES, which they bundled together in a fairly arbitrary way:
Next, each of 2407 European subjects was assigned into one of 10 groups based on geographic region: South:Italy, Swiss-Italian; Southeast: Albania, Bosnia-Herzegovina, Bulgaria, Croatia, Greece, Kosovo, Macedonia, Romania, Serbia,Slovenia, Yugoslavia; Southwest: Portugal, Spain; East: CzechRepublic, Hungary; East-Southeast: Cyprus, Turkey; Central:Austria, Germany, Netherlands, Swiss-German; West: Belgium,France, Swiss-French, Switzerland; North: Denmark, Norway,Sweden; Northeast: Finland, Latvia, Poland, Russia, Ukraine;Northwest: Ireland, Scotland, UK.
I don't know exactly why Switzerland should be bundled with Belgium, while Austria with Netherlands, or that Finns would be bundled with Poles, or Albanians, Greeks, and Slavs would be bundled in a broad "Southeast" group. Anyway, the authors don't use this POPRES sample much in their actual paper, although they present some results in the supplement, so I won't dwell on it further.
UPDATE II (Jun 4)
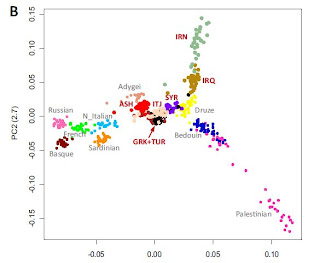
On the left we have panel B from Figure 1 in the paper, which shows the first two principal components in a regional context. Capitalized labels represent Jewish groups. Note that Iranian and Iraqi Jews don't show a particular relationship to Arabs (Bedouins and Palestinians). It would be interesting to see if there is a relationship with Iraqis or Iranians, which might indicate whether admixture with these local groups is responsible for Iranian/Iraqi Jewish distinctiveness. As in previous studies, most other Jews, including Syrian Jews, are located between Europeans and Druze; the lack of non-Jewish Euranatolian populations is especially baffling.
I am particularly interested in the seemingly very close relationship between Greek, Turkish, and Italian Jews. The GRK-TUR relationship is not that puzzling, as these are mostly Ottoman Jews who found themselves on different sides of national borders, and we would not expect them to be any different. But, why would they be so similar to Italian Jews? Speaking of Greek Jews, how many of them are of Romaniote and how many of Sephardic extraction?
UPDATE III (Jun 4)

The STRUCTURE analysis is also quite interesting. Jews seem to lack appreciable levels of East Asian (orange) or Sub-Saharan African (yellow) admixture, or of Central/South Asian admixture (green). The lack of E/C/S Asian admixture is especially damning of the Khazar hypothesis.
We should probably not interpret the three main visible components ("European" blue, "Mozabite" purple, "Near Eastern" pink) as representing ancestral proportions of European, North African, and Near Eastern elements. For example, Mongoloids have some "purple" while it is unlikely that they have North African admixture; so, while purple has an obvious relationship to Mozabites, it is not a good fit for an ancestral population group. Its substantial presence in the Near East also precludes such an easy interpretation.
Nor can we easily infer the percentage of "European" and "Near Eastern" admixture in Jews. The "Pink" element seems to grade from prominence among Iranian Jews to insignificance among Basques, but what did the original European and Jewish groups look like? Depending on how close they were to the Basque and Iranian Jewish end of the gradient, quite different admixture proportions would arise.
In more mathematical terms, a gradient can be represented as a single variable x going from 0 to 1, e.g., pink/(pink+blue) in the STRUCTURE analysis or relative position between Basques and Druze in the PCA figure above. But x can be expressed in an infinite number of ways as a weighted summation of two other numbers between 0 and 1. If ancestral groups were
exactly like Basques and Druze or they were
exactly pure blue and pure pink, then we could arrive at exact ancestral proportions for living Jews, but unfortunately, unlike situations where
clear-cut well-differentiated ancestral g
roups exist to act as yardsticks, this does not appear -as of yet- to be the case for intra-Caucasoid variation.
UPDATE IV (Jun 4)
From the paper:
Admixture with local populations, including Khazars and Slavs, may have occurred subsequently during the 1000 year (2nd millennium) history of the European Jews. Based on analysis of Y chromosomal polymorphisms, Hammer estimated that the rate might have been as high as 0.5% per generation or 12.5% cumulatively (a figure derived from Motulsky), although this calculation might have underestimated the influx of European Y chromosomes during the initial formation of European Jewry. Notably, up to 50% of Ashkenazi Jewish Y chromosomal haplogroups (E3b, G, J1, and Q) are of Middle Eastern origin,15 whereas the other prevalent haplogroups (J2, R1a1, R1b) may be representative of the early European admixture. The 7.5% prevalence of the R1a1 haplogroup among Ashkenazi Jews has been interpreted as a possible marker for Slavic or Khazar admixture because this haplogroup is very common among Ukrainians (where it was thought to have originated), Russians, and Sorbs, as well as among Central Asian populations, although the admixture may have occurred with Ukrainians, Poles, or Russians, rather than Khazars. In support of the ancestry observations reported in the current study, the major distinguishing feature between Ashkenazi and Middle Eastern Jewish Y chromosomes was the absence of European haplogroups in Middle Eastern Jewish populations.
I would not be so quick to assign haplogroups to European or Middle Eastern origin. For example, G seems to have originated in the Middle East, but it is quite plentiful in substantial parts of Europe. So, while its ultimate origins may be West Asian (it arose in a man who lived in West Asia thousands of years ago), its proximate origin may be European in some particular case.
As I have argued before, I doubt E3b (or E1b1b) was an original Jewish lineage, J2 probably represents Iranian/Euranatolian admixture in Jews, while J1 (or a subset thereof) has strong Semitic connotations.
UPDATE V (Jun 10):
AJHG doi:10.1016/j.ajhg.2010.04.015
Abraham's Children in the Genome Era: Major Jewish Diaspora Populations Comprise Distinct Genetic Clusters with Shared Middle Eastern Ancestry
Gil Atzmon et al.
Abstract
For more than a century, Jews and non-Jews alike have tried to define the relatedness of contemporary Jewish people. Previous genetic studies of blood group and serum markers suggested that Jewish groups had Middle Eastern origin with greater genetic similarity between paired Jewish populations. However, these and successor studies of monoallelic Y chromosomal and mitochondrial genetic markers did not resolve the issues of within and between-group Jewish genetic identity. Here, genome-wide analysis of seven Jewish groups (Iranian, Iraqi, Syrian, Italian, Turkish, Greek, and Ashkenazi) and comparison with non-Jewish groups demonstrated distinctive Jewish population clusters, each with shared Middle Eastern ancestry, proximity to contemporary Middle Eastern populations, and variable degrees of European and North African admixture. Two major groups were identified by principal component, phylogenetic, and identity by descent (IBD) analysis: Middle Eastern Jews and European/Syrian Jews. The IBD segment sharing and the proximity of European Jews to each other and to southern European populations suggested similar origins for European Jewry and refuted large-scale genetic contributions of Central and Eastern European and Slavic populations to the formation of Ashkenazi Jewry. Rapid decay of IBD in Ashkenazi Jewish genomes was consistent with a severe bottleneck followed by large expansion, such as occurred with the so-called demographic miracle of population expansion from 50,000 people at the beginning of the 15th century to 5,000,000 people at the beginning of the 19th century. Thus, this study demonstrates that European/Syrian and Middle Eastern Jews represent a series of geographical isolates or clusters woven together by shared IBD genetic threads.
Link



111 comments:
It seems to me that this paper demonstrates what I have said in the past, and the Italian component is recognized and I think massive: many Y haplogroups, beginning from the most ancient R1b1*,R1b1b2, R1b1b2a and the mithocondrial K, which is, from my K1a1b1/9932A, certainly Italian.
Very interesting, thanks. In this I think we are pretty much in agreement.
I'd love to see how they compare with Turks and Greeks in particular. I imagine that the paper will be freely available in six months.
The Northeast set is basically made up of Poles and a few Russians from Moscow.
There is only one Finn, as well as one Latvian and one Ukrainian.
2 comments:
1. To add to your wish list of things that would have been good to have: Sephardi data.
2. Your comments on E1b1b/E3b seem quickly written and could mislead people. I think the most important point is that there are different clades. E-M123 is very likely to be quite old amongst Levantine populations. E-V12 and E-V22, perhaps also. E-V13 may well have originated in the Levant, but I believe the evidence strongly suggests that nearly all surviving E-V13 dispersed from Europe. Did any make it back to the Levant in time to be "original Jewish"? It certainly could have, but I think the neutral hypothesis is that at least most E-V13 in Jewish populations entered the Jewish population outside of the Levant. In summary: E-V13's connection to Jewish ethnohistory has so many options possible that we can not really say much.
(Which in the end agrees with your doubts about what the authors say.)
"I doubt E3b (or E1b1b) was an original Jewish lineage"
An E3b component of the original Jewish mix makes plenty of sense to me. Both J1 is the modal Arab pastoralist haplotype, while E3b is the modal Berber haplotype. The descendants of Arab pastoralists have a significant minority of E3b. It is entirely reasonable to link that there would be some admixture of adjacent populations speaking languages from the same language family with similar lifestyles from ancient times, when pastoralism was first present in both areas ca. 4500 BCE or earlier. One sees similar fusion of adjacent pastoral cultures taking place now between Hasua and Fulani peoples of the Sahel.
If Jews are, as all evidence seems to indicate, one of the descendants of Semitic pastoralists in the Near East and developed a truly distinct identity from them only 1900 BC to 1300 BCE, the Jews would acquire the E3b as part of the Near Eastern Semitic pastoralist gene pool from which they emerged and grew distinct.
The fact that it is present in similar proportions in the Druze and Palestinians, and in higher proportions in contemporary Beudoins supports the idea that E3b predates Jewish ethnogenesis, and ceased to flow into their combined gene pools when all of these groups transitions from a nomadic to a non-nomadic lifestyle, while the Beudoins, who continued to have a nomadic lifestyle saw continued infusions of E3b into their gene pool.
"I am particularly interested in the seemingly very close relationship between Greek, Turkish, and Italian Jews. The GRK-TUR relationship is not that puzzling, as these are mostly Ottoman Jews who found themselves on different sides of national borders, and we would not expect them to be any different. But, why would they be so similar to Italian Jews?"
I also don't see why this is surprising. Greek and Turkey and Rome were all core urbanized areas of the Roman Empire at the time of the fall of the Temple in 70 AD when the Jewish diaspora commenced. Refugees then and now go where the opportunities are, and in 70 AD that would have been in the urban centers of the Roman empire. Also, reasonably large and isolated Jewish immigrant communities in Roman empire Jewish centers (something we know existed from the historical record) would have been better able to completely avoid admixture with other populations than smaller and more isolated Jewish immigrant populations in the relatively rural hinterlands of Central and Eastern Europe.
Iraqi and Iranian Jews were clearly spread even thinner in even smaller communities and hence had to accept even greater admixture.
@Onur
The Adygei in the PC chart is a non-Jewish Caucasian population that speaks a North Caucuasian language.
Given the apparent historical origin of Caucasian Jews from Iran, one would expect the Caucasian Jews to cluster somewhere between the Adygei and the Iranian Jews.
The Caucuasian Jewish population was apparently small (about 25,000) from 1926-1959 and there was also apparently a lot of interclan marriage with Chechens. Presumably their population was even smaller in earlier times. The population has exploded since 1959 to about 250,000 today and a majority now live in Israel, where there has probably been great admixture with other Jews in the last half century.
I suppose that your interest is that Caucusian Jews would be one way to explain the anomolously high proportions of J1 in Dagestani populations from a recent (i.e. post-Roman Empire) rather than an ancient migration.
Thank you for this very interesting study, please would you answer to these 3 question:
1/(At K=6)Is the 10-20% European (blue) component amongst Arabs and Mozabites due to historical European genetical influx (ie crusades and colonisation)?
(Keeping in mind that Mozabites were not a target of crusades and their endogamy and geographical isolation in Sahra should had prevented them from colonial admixture?)
btw why the middle eastern (pink) component is lacking amongst them , knowing that they are most likely the result of neolithic norafrasan migrations from middle-east-Egypt included-as it's shown by their Y-DNA and to a lesser extent by their mt-DNA?
2/Also , I thought that Europe was populated by hunter-gatherers and -after the neolthic revolution in the region of fertile crescent and the consequent demographic explosion-by farmers who migrated there from the fertile crescent; so why the middle eastern (pink) component is so low amongst Europeans? (even lower than the north African [purple] component for the Sardinians!)
3/And could we conclude that, thus, blondism and other caucasian features amongst middle-easterners are the result of this historical European influx (even if very ancient depictions of some middle easterners types contradict this)?
Thank you for your attention.
An E3b component of the original Jewish mix makes plenty of sense to me. Both J1 is the modal Arab pastoralist haplotype, while E3b is the modal Berber haplotype. The descendants of Arab pastoralists have a significant minority of E3b. It is entirely reasonable to link that there would be some admixture of adjacent populations speaking languages from the same language family with similar lifestyles from ancient times, when pastoralism was first present in both areas ca. 4500 BCE or earlier.
First of all, Berbers have a particular type of E1b1b, E-M81. It's not clear how important that is among Jews; I'm sure there are data about it somewhere, so feel free to chime in.
Second, Berbers were not adjacent to Jews. In fact Jews are not even mentioned in classical sources until Hellenistic times, so the overall impresion is clearly of a fairly small people that didn't have substantial intercourse with its Mediterranean neighbors.
I also don't see why this is surprising. Greek and Turkey and Rome were all core urbanized areas of the Roman Empire at the time of the fall of the Temple in 70 AD when the Jewish diaspora commenced. Refugees then and now go where the opportunities are, and in 70 AD that would have been in the urban centers of the Roman empire
The authors claim that 2.5ky created the differences observed between Iranian and European Jews. If Italian and Euranatolian Jews are descended from the 70AD diaspora, then we'd expect a divergence of similar magnitude, which is NOT what we see.
One needs to differentiate between jews and "Semitic" groups encompassing people and language. Jews are simply a subset of Semitic speakers and were likely several haplogroups before being dispersed throughout Europe and Western Asia. It's a known fact that J2 peaks along the fertile crescent where the Amorite tribes dominated the Sumerians. I don't see how one could rule out J2 as being Semitic...IMHO it seems like it would have been among the Amorite speakers.
Andrew said,
"The fact that [E3b] is present in similar proportions in the Druze and Palestinians, and in higher proportions in contemporary Beudoins supports the idea that E3b predates Jewish ethnogenesis, and ceased to flow into their combined gene pools when all of these groups transitions from a nomadic to a non-nomadic lifestyle, while the Beudoins, who continued to have a nomadic lifestyle saw continued infusions of E3b into their gene pool."
Where are the data to support this paragraph? As I recall, Nebel et al. have found haplogroup E-SRY4064 to be most common in Ashkenazi Jews and second-least common in Bedouins (after Kurdish Jews) among all sampled populations of modern Israel. Haplogroup E3b generally is rather uncommon in populations of Arabia, so I think the neutral expectation is that E3b should not occur with particularly high frequency among Bedouins. E3b actually is significantly more common among settled populations of the Nile Valley than it is among recently nomadic or semi-nomadic populations of the Arabian Peninsula.
Thanks for posting the graphs Dienekes. I had understood they compared Jews with Turks and Greeks, which I understand it is important in understanding the diaspora and the true Jewish genetic origins. Sadly this doesn't seem to be the case. :(
Still they do look like West Asian but not strictly Palestinian, because they lack the small but noticeable African affinity that must be very old in the southern West Asian region.
Again the PC graph places them roughly where Turks and Greeks should be if sampled, again suggesting an Anatolian origin, in agreement with the historical data.
"The authors claim that 2.5ky created the differences observed between Iranian and European Jews. If Italian and Euranatolian Jews are descended from the 70AD diaspora, then we'd expect a divergence of similar magnitude, which is NOT what we see."
This is the million dollar question, what happened historically in those areas 500 years b.c.e?
I also thought, that their would be a genetic trail coming out Iraq, as what I've read, the orignal Hebrews came out of Sumeria, from the city of Nippur. Did they disappear or get overwhelmed genetically by another middle eastern core group?
Wouldn't the Adyghe people be the nearest to the historic Khazars? In the charts the Adyghe seem very close to the Ashkenazi...
"The authors claim that 2.5ky created the differences observed between Iranian and European Jews. If Italian and Euranatolian Jews are descended from the 70AD diaspora, then we'd expect a divergence of similar magnitude, which is NOT what we see."
So, it appears that Iranians were isolated from European Jews, probably more or less completely (a 2,500 year old separation would suggest a pre-diasporan separation for them, and the date would correspond roughly to Babylonian exile).
How could the separation of other Jews be less than a diasporan 1900 years? There was regular trade between Greece, Ottoman Turkey and Italy, and to a less but real extent also with Europe, more or less continuously after the diaspora, in which Jews were historically documented major participants (e.g. they were prominent traders in the Kingdom of Venice for its entire existence) and there is also historical documentation for dissemination of Talumdic scholarship all across this network. So, perhaps this trade also included arranged marriages (they must have been concerned about inbreeding in small endogamous communities given religous rules on incest) and perhaps this was sufficient that these populations didn't totally diverge after the diaspora.
"Second, Berbers were not adjacent to Jews. In fact Jews are not even mentioned in classical sources until Hellenistic times"
My argument is that indigeneous North African-Proto-Semite admixture preceded Jewish ethnogenesis taking place from roughly 4500 BCE through the Hyskos Dynasty in Egypt. The fact that the level of admixture is similar for the Near Eastern Jews, Palestinians and Druze suggests that admixture preceded Jewish ethnogenesis.
I was overspecific use "Berbers" when I meant indigeneous North Africans among all of whom E1b1b1 is common. The Berber subgroup of North Africans is largely E-M81 which is rare in other groups except Iberians including Sephardic Jews. E1b1b1 haplotypes broke up into subtypes around the time of the Neolithic revolution.
North Africans with E1b1b1 were separated from the Semitic nomads of the Arabian Penninsula with J only by the Nile Delta and largely uninhabited territory. They acquired domesticated animals from the Near East, so they were in communication with their neighbors. Proto-Semites would have been even closer during the Hyskos regime in Egypt (which is when most historians think Jews made their way to Egypt prior to Exodus), and also would have had exposure via Old Kingdom Egyptian rule of the Levant.
"Haplogroup E3b generally is rather uncommon in populations of Arabia . . ."
E3b and Mozabite contributions generally are common to the same extent among the Levantine populations studied in the linked paper (Jews, Druze, Palestinians). There are more Mozabite contributions are more common in Bedouins than they are in Levatine Arab/Druze populations suggesting a longer period of admixture.
E3b is present at significant frequencies in Arabia in two of its subtypes.
Contrary to Dienekes, I see some E3b haplotypes and J2 as clearly part of the original gene pool of the Jews at the time of ethnogenesis, through admixture of proto-Semites prior to Jewish ethnogenesis. E-M123 (i.e. E1b1b1c) is found at high percentages in both Ashkenazi and Sephardic Jews making a strong case that it is part of the original Jewish genetic mix.
E-M81 (i.e. E1b1b1b) associated with Northwest Africa and Iberia and found most commonly in Sephardic Jews, may not be part of the original gene pool of Jews, and instead be a product of later contact with North Africans probably in the Islamic era. But, the basal E1b1b1*/M35* subclade is found in 20% of the Ashkenazi Jewish population, suggesting that it is a part of the founding population of Jews.
The notion that J2 is a recent admixture also doesn't ring true, since it is found in Ashkenazi Jews 24%, Palestinian Arabs 16.8% and Sephardic Jews, 15.4%. J2 is the Cohen Model Haplotype for cripes sake.
Andrew Oh-Willeke said...
"An E3b component of the original Jewish mix makes plenty of sense to me. Both J1 is the modal Arab pastoralist haplotype, while E3b is the modal Berber haplotype. The descendants of Arab pastoralists have a significant minority of E3b."
I can not see the relevance of J1 or Berbers in this story. Most E1b1b (E3b) amongst Arab pastoralists is either E-M123 or E-V22, both of which are also found in Jewish populations. E-V13, the type of E1b1b which is most common in European Jews is not very common outside of Europe, and is common amongst non Jewish Europeans. Having said that it DOES appear in areas like Lebanon. So there IS a chance that V13 was in the Levant from early times.
Dienekes said...
"First of all, Berbers have a particular type of E1b1b, E-M81. It's not clear how important that is among Jews; I'm sure there are data about it somewhere, so feel free to chime in."
Every study I can recall seeing shows low E-M81 in Jewish populations, and it is pretty much absent in the Middle East as a whole, apart from in cosmopolitan areas where it can be found in trace amounts (probably brought to Turkey and Cyprus by Sephardim?)
Intriguingly, the only place where M81 looks native outside of the Maghreb is perhaps the Horn of Africa. In summary all signs are that M81 started in one part of Africa and spread to another part of Africa, and has nothing to do with the Middle East.
Anyone looking for more data can search by country or region name on the E-M35 haplowiki: http://www.haplozone.net/wiki/
"First, thank you to everyone who kindly forwarded a copy of this study. I greatly appreciate it.
It appears that there is a significant misunderstanding about the findings of this study.
The study neither refutes nor supports the hypothesis that Ashkenazim, particularly those of Eastern European descent, descend in some part from an ancient Khazarian source. On that subject, the authors state: "Admixture with local populations, including Khazars and Slavs, may have occurred subsequently during the 1000 years (2nd millennium) history of the European Jews." The authors then cite to Hammer's study on the R1a Levites as supporting this hypothesis. Interestingly, the authors suggest that Q may be derived from a Middle Eastern source among Askhenazim, but J2 may be derived from a European source. Other Y haplogroups among Ashkenazim with possible European origins are R1b & R1a1 (no surprise there).
To distill down the other findings, basically the authors found that Jews comprise a loose genetic cluster, with Ashkenazim falling as something of an outlier of the group. The authors suggest that this is due to a high rate of admixture among Ashkenazim with their non-Jewish European neighbors, reaching a rate of 30-60%, as well as a dramatic bottleneck effect. On the other hand, the loose clustering suggests a shared ancestry reaching back to an ancient Israelite past.
Interestingly, the study found close genetic ties between Ashkenazim and Northern Italians, French and Sardinians (as well as Syrian Jews). The authors suggest the genetic affinity "favors the idea of non-Semitic Mediterranean ancestry in the formation of the European/Syrian Jewish groups." The authors suggest that the core Ashkenazim population was formed from a mixture of Jews who migrated or were expelled from Israel, with those who converted during either Hellenic-Hasmonean or Greco-Roman times.
One intriguing findings is the apparent genetic connection between the Adygei, a Caucasian population, and Ashkenazim. This is not the first study to have found this unexpected genetic link. Sorry, can't recall the title of the previous study, but it found genetic relatedness between the Adygei, Ashkenazim, Egyptians and Greeks. The suggestion was that these groups might contain genetic remnants of a very ancient Mediterranean population that pre-dated the Arab populations that settled later in the Middle East.
A few other interesting findings: The Iranian & Iraqi Jews form a distinctive cluster and show the least "European" admixture (and a lot of inbreeding). The split from the rest of the Jewish groups is estimated at 2500 years ago.
Also, the Sephardic, Italian Jewish and Syrian Jews showed "a low level component (8-11%) that was shared with the North African Mozabite population."
Ellen Coffman (forwarded by Gioiello)
"J2 is the Cohen Model Haplotype for cripes sake".
There are two CMHs: one for J1 and one for J2. While it's maybe true that ancient Jews had some J2, the apportions should be more like modern Palestinians, what rings true in the Sephardi case but not in the Ashkenazi case. Also they should have much higher apportions of J1 in both cases, I think.
"Interestingly, the authors suggest that Q may be derived from a Middle Eastern source among Askhenazim, but J2 may be derived from a European source".
That IS INTERESTING, thanks Gioello.
"On the other hand, the loose clustering suggests a shared ancestry reaching back to an ancient Israelite past".
I've been thinking about that PC graph and, while it may be an effect of the sampling strategy, Euro-Turco-Syrian Jews do not cluster more tightly than, say, Basque and Russians (a most unlikely cluster) and Ashkenazi do not cluster more tightly than, say, French. It is true that they do show clustering in the K-means analysis but including also all other West Asians, what is not too clarifying.
"The authors suggest that the core Ashkenazim population was formed from a mixture of Jews who migrated or were expelled from Israel, with those who converted during either Hellenic-Hasmonean or Greco-Roman times".
Bravo! It probably applies also to other Jews, specially those who cluster close to Ashkenazi: the Euro-Turco-Syrian group.
"One intriguing findings is the apparent genetic connection between the Adygei, a Caucasian population, and Ashkenazim".
The link is probably emphasized because there are no other Aegean-Pontic populations to compare with. They always use Adygei for the same reason they use Mozabites: it's an easily available sample. IMO the Adigey connection represents an Anatolian connection in fact.
"Wouldn't the Adyghe people be the nearest to the historic Khazars?"
Maybe or maybe not. They are not a Turkic population. Russians should represent Khazars at least as well.
IMO they represent the Pontic-Aegean gene pool that fails once and again to be compared with. Fear of finding the truth? Notice that all these studies on Jewish genetics are made by Jews, who in turn tend to have an ideological bias and a tendency to idealize their origins. The last thing they want to know is that they are not from Palestine but from Turkey: it would be a serious blow to the Zionist ideology.
The only paper I know so far that compared Jews (Ashkenazi) with Pontic-Aegean peoples is Bauchet 2007, which was not focused on Jewish but European ancestry. In it Jews clustered very very tightly with Greeks and Armenians, while also showing some other European ancestry.
The study on Jews with reference to Middle Eastern groups and Europeans shows me that what was said about Jews can just as easily be said about Europeans: Europeans have Middle Eastern and North African admixture just like the Jews and like the Middle Easterners. Their results are very similar as far as the three main players represented by ancestral blue, pink and purple.
So what exactly is the difference between these Jews and the Europeans and the Middle Easterners?
In the PCA map why are the Russians placed away from the Adygei? In most studies using Russians, albeit North Russians, the Russians are located very close to the Adygei. From the frappe results it would seen that North Russians were used for the study, so why the different placement?
It is also about time you all dropped using haplogroups as being of anything significant other than a population statistic of little importance.
I am commenting from memory so this is only a general comment, but from what I recall, there are historically very strong ties between Greek and Turkish(Ottoman) Jews, and Italian Jews. Following the expulsion from Spain and Portugal, Livorno for example, became a refuge, and a large trading center with the Ottoman empire. Initially, some separation by origin was maintained among the different Jewish groups, in the trading centers of the Empire and in North Africa, but as I remember, that eventually broke down. The connection between Livorno, Tuscan Jews and Salonika persisted into the period right before the second world war, with famous rabbis who ultimately settled in Salonika, bearing names from the Tuscan countryside.
"In the PCA map why are the Russians placed away from the Adygei? In most studies using Russians, albeit North Russians, the Russians are located very close to the Adygei. From the frappe results it would seen that North Russians were used for the study, so why the different placement?"
Because PC analysis is easily distorted by sample size. In this case, as in others, the Jewish sample is so big that most of the structure is determined only by the internal structure of Jews themselves, strongly distorting the rest. As Adygei cluster well with Jews they appear at the center of the graph, while Basques and Russians (in the European sample) seem to be the most distant to Jews in general and hence pop up at one extreme and oddly close to each other. What brings them together is surely nothing else but lack of Jewish or otherwise West Asian component.
That condition does not apply to the Adigey and probably would not apply, at least not so extremely, to more southernly Russians either.
In essence the PC1 is an Europe-West Asia axis, while the PC2 is a Palestinian-Iranian Jewish (tending towards regular Iranians?) axis, in which all Europeans are essentially neutral.
Most Jews, and the Druze too, are pretty much neutral to both components, indicating that they are intermediate between the most characteristic European and West Asian populations and also between Iranian Jews and Palestinians.
This lack of specificty for the bulk of such a large sample probably just means hybridation and/or intermediate geographic origin as the Anatolian one I keep proposing.
@Onur:
"... am I right in my inference?"
That's what I think, yes.
I strongly suspect a differential founder effect among Diaspora Jews at their origins, which had as main center Asia Minor (and to lesser extent Syria and other localities), where they must have assimilated lots of local natives because at that time Judaism was very much proselytist (something that changed when Christianity and Islam forced them to become non-proselytist in order to tolerate them).
Sadly this hypothesis is systematically not being tested, with Turks and nearby populations not being sampled and compared. The closest proxy in this study are probably Adigey and surely Druzes too and they seem to confirm my suspicions. But the final word will only come when the appropriate populations are effectively compared, in particular Turks.
"The suggestion was that these groups might contain genetic remnants of a very ancient Mediterranean population that pre-dated the Arab populations that settled later in the Middle East."
-- I think she's on the right track with this point. However, then this throws into question the YDNA line of Abraham, which is basically what we are discussing. Otherwise, the jewish people would be a combination of several indigenous groups inhabiting from Iraq to the Near East and everywhere inbetween. Realistically, the latter situation seems more probable.
Let's not forget that Abraham is a mythical figure, and stories about him should be treated as such - there is no evidence that he actually existed.
"I think she's on the right track with this point".
That's what you want to believe, that's what all Jewish nationalists want to believe. But the only group that clusters with Jews in the Levant are Druze who are also historical immigrants with an essentially Anatolian origin.
As all these research papers are made by Jews with a clear Jewish-centric bias, key evidence (Anatolians and other northern West Asians) is being skipped once and again in the same systematic pattern that can only raise our eyebrows and our skepticism.
Arabs (a bunch of nomads from the desert originally) could not alter the genetic landscape in any meaningful manner. The Arab replacement hypothesis has been dismantled once and again, not just in Palestine but also in North Africa.
The origin of Druzes was discussed in depth at this blog in 2008.
There's no evidence that they are Levantine. In fact they, like most Jews, cluster best with the Aegeo-Pontic peoples all the time. They look anything but Levantine, though they may have more Egyptian ancestry share than Jews.
"That is not entirely true, as a sample of 4 Turks was included in this study; they did not look particularly close to Jews".
Really? As I have no access to the paper I'm missing info.
"I doubt that e.g., Turks or Iranians are ready to jump at the opportunity to support Israeli research".
Until this past week Turkey was the strongest ally of Israel in West Asia (shared anti-Arab stand) and I doubt that Israeli or US researchers would have any problem whatsoever sampling Turks, Kurds, Armenians or Georgians.
Yet they do not. Why? Because of pre-conceptions or even maybe post-conceptions, as we can't be sure they did not do it, found something and then hid it under the rug.
"Then, if I understood you correctly, according to you there should be no pure (relatively speaking) Jewish communities in the world today"
And presumably never have been. Jews are just a grouop of people with a common religion, exactly the same as is any other religious group. Their myths about a common ancestry from a single man are just that: myths. It's just that the Jewish stud book has been relatively closed for the last 1500 years. Until then, as Maju said, 'Judaism was very much proselytist (something that changed when Christianity and Islam forced them to become non-proselytist in order to tolerate them)'.
"the jewish people would be a combination of several indigenous groups inhabiting from Iraq to the Near East and everywhere inbetween".
Exactly.
I don't have a copy of the paper in question, but I'm not sure I follow why J2 is being ascribed to "early European admixture" unless the "early" here refers to events prior or during the Jewish ethnogenesis.
Like I said, I don't have this study, but
1) in some other studies, Sephardic Jews have a higher proportion of J2 than J1 (and a higher proportion of J2 compared to Ashkenazi samples)
2) Samaritans notably have a higher proportion of J2 compared to other haplogroups
3) as someone mentioned, there is the J2 CMH which the recent CMH study has shown to be old
4) J2 is predominant amongst the coastal Lebanese Maronites, which Zalloua concluded was the modal haplogroup of the Canaanites.
For obvious historical reasons Palestinian Arabs (PA) aren't going to be a suitable comparison group:
1) Documented history of Arab-Islamic expansion and settlement in Palestine
2) PA tribal genealogy connecting them to old Yemeni, Hijazi, and Nejdi Arabian tribes and personages
3) the Galilee Modal Haplotype presence in the PA samples (and its absence in Jewish samples)
4) the fact that the CMH (neither J1 nor J2) does not occur in any statistically significant frequency in PA samples
5) the significant incidence of the Bedouin Modal Haplotype in PA samples (and in general, the close clustering of PA to Bedouin and Arabian Peninsula samples)
6) the low haplotype diversity of the PA samples, indicative of a more recent "founding event" (Arab-Islamic expansion)
I think there have been enough studies that have connected J2 with the fertile crescent (including the levant) long before the Hellenic or Roman periods
As for the Druze, I would point out that though they speak Arabic, they don't consider themselves to be Arabs.
They are Arabs whether they consider themselves to be or not. As to what their origins are, I would say that they've probably had less time and opportunity to mix with other Arabs as they were formed fairly early after the Islamic conquest, so it's reasonable that they are genetically less Arab-ized than some other Levantine groups.
The origin of Druzes was discussed in depth at this blog in 2008.
Absolutely no evidence in that discussion that Druze are of "Anatolian origin".
"Arabs (a bunch of nomads from the desert originally) could not alter the genetic landscape in any meaningful manner. The Arab replacement hypothesis has been dismantled once and again, not just in Palestine but also in North Africa."
Dont forget that Canaanites, Arameans, Hebrews, Akkadians, Sabaians, Amorites and other Arabians too were Desert nomads.(but perhaps before this were farmers as agriculture terms in Arabianic languages are very old and has Arabianic etymologies and also ie+dravidian+turkic counterparts as in Akkadian akar/English acker/Turkish eker)
As for the replacement hypothesis, I dont think so as "J1" is solidly present in north Africa, Sudan, Abyssinia, Levant and mesopotamia.
Dont forget that polygamous (4 wives+X number of islamic concubines) muslim Arabs could have, very well and shortly, submerged local Arabians.
More I read more I think that (at least prior to the 20 th century), all Arab speaking folks descend largely from Arabs (except Egypt, but in this case the Egyptian case is perhaps similar to Ibero-Aquitanian speaking west Europeans adopting ie language).
"As for the replacement hypothesis, I dont think so as "J1" is solidly present in north Africa, Sudan, Abyssinia, Levant and mesopotamia."
-- Right..., so in your eyes language replacement only works one way -- if you are a West European Ibero-Aquitanian. One thing to note is the "Middle-Eastern" ancestry in the STRUCTURE bar graph peaks at the heart of the Sumerian and Elamite nations. However, according to Sumer inscriptions the Semites actually came from further west. So I ask which is it? There is language replacement somewhere...
"PA tribal genealogy connecting them to old Yemeni, Hijazi, and Nejdi Arabian tribes and personages"
I'd like to know the exact extent of such genealogies: how many Palestinians actually claim to have such ancestors by pure patrilineage?
We know that Yemeni J1 is very low in diversity, while Palestinian and North African one is instead very diverse. This suggests that J1 expanded from North (Palestine) to South (Arabia), what surely happened within the frame of the Neolithic. Other research on Arabian DNA (Abu Amero for example) also support such pattern of settlement of the peninsula, eventually reaching Yemen.
"3) the Galilee Modal Haplotype presence in the PA samples (and its absence in Jewish samples)
4) the fact that the CMH (neither J1 nor J2) does not occur in any statistically significant frequency in PA samples"
This is supportive of a distinct origin, at least to some extent, of the two ethnicities but it is not demonstrative as such of who are the most genuine descendants of ancient Jews, if any. The CMH anyhow does occur among Palestinians, though it's logical to understand that most Cohanim remained within the Jewish faith, right?
"5) the significant incidence of the Bedouin Modal Haplotype in PA samples (and in general, the close clustering of PA to Bedouin and Arabian Peninsula samples)"
Almost invariably, Bedouins used in genetic research refers to Negev Bedouins, which are a very specific population with clear links (and possibly deep roots) in that part of Palestine.
Anyhow, as said before, peninsular Arabs should have Neolithic ascendance from Palestine and I'd dare say that most of it has that ultimate origin. The opposite is surely possible but less important overall probably. Bedouins do not normally settle as farmers or artisans, right?
"6) the low haplotype diversity of the PA samples, indicative of a more recent "founding event" (Arab-Islamic expansion)"
Source? That would be really interesting to analyze, though I doubt it has any reality to it.
Kidproquo:
The paper says:
"Notably, up to 50% of Ashkenazi Jewish Y chromosomal haplogroups (E3b, G, J1, and Q) are of Middle Eastern origin,15 whereas the other prevalent haplogroups (J2, R1a1, R1b) may be representative of the early European admixture."
And refers to Li et al. (2008). 'Worldwide human relationships inferred from genome-wide patterns of variation'. Science 319, 1100–1104.
I imagine that there is haplotype evidence that it is closer to "European" J2 (J2b surely) than to Levantine one, however I wonder if it may also be Anatolian J2.
"there is the J2 CMH"
The true CMH is probably J1 (the most common by far among Kohanim themselves) with the J2 one being a red herring with all likelihood.
"J2 is predominant amongst the coastal Lebanese Maronites, which Zalloua concluded was the modal haplogroup of the Canaanites".
I'd take that with a tablespoon of salt, sincerely. Canaanite is of course a relevant proto-historical term but they also had their genesis, which must have been rooted, Semitic invasions apart, on the Neolithic populations of the area.
The reality is that J2 seems to have a more Northern distribution in West Asia than J1 overall and this probably reflects an early Upper Paleolithic (or at most early Neolithic) distribution pattern. So talking of 'Canaanites' in this context is like the proverbial tree and the forest.
My opinion anyhow.
"For obvious historical reasons Palestinian Arabs (PA) aren't going to be a suitable comparison group:
1) Documented history of Arab-Islamic expansion and settlement in Palestine".
Settlement? Can you refer to such documents?
IMO This is the usual hasbara confusionist blank statement about Palestine. The latest date possible for a massive resettlement of Palestine would be after the Roman-executed genocide. Remember that Arabia is essentially a desert and that Arabs expanded as aristocratic conquerors and religious missionaries, not as settlers.
@Adyghe: I have already replied to you by email but I'd like to add that disguising Jewish supremacism and racism under victimism and unfounded accusations of Judeophobia on my side is lowly. Respect and you shall be respected.
@Dienekes: If you read my last post in that very interesting discussion of 2008 you can see:
Liran Shlush (quoted): "Actually figure 2 can answer many of the questions you have raised regarding Palestinians. As you can see Palestinians have low migration rates with Druze and high migration rates with other Near eatern populations. This can be true or a false positive results due to the different sampling methodologies".
Me: "Yes, figure 2 (that I overlooked before) is interesting.
"Following that figure the smallest divergence rate of Galilean Druzes is with Greeks and Adygeis (less than 2000 years). If the model is correct, then Druzes are derived from Caucasian Greeks (or something like that). The next divergence figure is with Egyptians (c. 2000 years). So what does that tell us about Druzes? If anything that they do not look Levantine at all".
'Caucasian Greeks' was obviously an approximating concept, of course. It can perfectly mean Anatolians and I'm pretty sure that was what I had in mind.
Evidence is lacking of course but it's not my fault but that of some geneticists who are clearly avoiding the issue. :)
@Gioello: I agree with much of what you say but:
"Dont forget that polygamous (4 wives+X number of islamic concubines) muslim Arabs could have".
Polygamy, even among Muslims, can only be a rare occurrence, a privilege of the wealthy and powerful. The common man will probably just have one wife, naturally. And anyhow, women also transmit their genetics, believe me. ;)
"If that had been case then Arabization would have happened in non-Semitic areas like Iran & Anatolia"
Anatolia was never conquered by the Arabs: the reached faster to France than into Asia Minor. There was and still is Arabization everywhere in the early Caliphate area, as well as in later Arab-conquered areas, like Sudan. Iran is in fact the only meaningful exception, probably because of one single reason: the prestige of Iranian culture that in fact was more influential in the new Islamic cultural region than the competing Greco-Roman one (though the hijab has a Greek origin AFAIK). In a sense the Arab Caliphate became the last Persian empire and moving the capital to Baghdad was also meaningful of that "Iranizing" trend.
"But the conclusions being reached are very convenient (and daresay it Orientalist); basically all the non-Muslims and "heretic Muslims" (Druze) population are autochthonous Middle Easterners but the Muslims are "South Arabs" converts".
Agreed. It's totally arbitrary. In the beginning of the Caliphate there were few Muslims in all conquered provinces and there was no pressure for conversion (specially because Muslims were exempt from taxes), later things changed and gradually most locals converted to Islam.
Anyhow, Semitic languages were the easiest ones to subsume into Arabic because of proximity. Similarly over here one Romance has replaced another through the centuries but replacing Basque has proven a tougher task (and vice versa: recovering Basque among Spanish speakers, while Catalan instead had that task easier because they are mutually intelligible languages). I imagine that Hebrew, Aramean and Chaldean speakers easily shifted to Arabic or a dialectal hybrid form.
Thank you mr Maju, for your very interesting comments.
I want to add that toponomy the neolithic population of the region of fertile crescent south of Taurus mounts was very likely Arabianic(=Semitic)or at least heavily Arabianic as:
-Just with the beginning of History we have an Arabianic culture/civilisation in middle Iraq with Arabianic toponomy (Babel=bab[gate]+el[god])=The Akkadians,this is the case also for the lower Euphrate valley and the southern area of coastal levant
-one of Hassuna,Half or Natufian cultures should be Arabianic or norafrasan or even afrasan speaking.
-The Arabianic etymology of the words for agriculture, domesticated animals, numbers which are in the same time shared by borrowing with proto indo-european (and some of them with Hurrian and Sumerian).
-Also in this same region existed 3 distinct Arabianic languages belonging to distinct branches (east Arabianic Akkadian, west Arabianic-but very close with modern Arabic-Ugaritic, and neither western nor eastern Arabianic Eblaitic.
As for the Jews, I think it's merely a religion (as Islam,Christianity,Hinduism etc...) not an ethny (I think "Hebri" should be THE ethny for the ones whose mother tongue is Hebrew).
But of course everyone could identify himself with the label he loves and live wherever he wants even if he do not descend from that ethny.
Of course the Israeli/Palestinian conflict will be resolved when Israel stop being an apartheid colonising country and will label itself as An Israeli nation where everyone whatsever his belief or mother tongue will be equal.
If Palestinians convert to Judaism and begin to call themselves Israelis (as Palestinian is a misnomer of a non Arabianic sea people) would this problem be resolved?(keeping in mind that Palestinians are genetically and haplotypically close to the Israeli Jews)
My question is this; is there a haplotype specifically associated with the Arab-Islamic conquests of the 7th century.
J1 seems to be the most obvious candidate but is there any more granularity than that?
To my mind the Arab-Islamic and Turko-Mongolian conquests were the two defining (unknown) demographic events of the past 1300 yrs.
part 2:
"The CMH anyhow does occur among the Palestinians"
The 6/6 J1-specific CMH occurs in singletons (not statistically significant) and the extended CMH's do not occur at all. If Muslim settlements did not alter the genetic landscape of Palestine one would expect a similar incidence of the CMH's in PA samples.
"it's logical to understand that most Cohanim remained within the Jewish faith"
Under what rule of logic do you make the inference that Cohanim remained within the Jewish faith but non-Cohanim didn't?
"Almost invariably, Bedouins used in genetic research refers to Negev Bedouins, which are a very specific population with clear links (and possibly deep roots) in that part of Palestine. "
Not to invoke an anthropological platitude, but nomads are more likely to eventually become settled populations than settled populations are likely to become nomads. And more so, Arab Muslim bedouin are more likely to become settled Arab Muslims than Rabbinical Jews are likely to become Arab Muslim bedouin.
At the end of the day, the genetic data regarding the significant proportion of J2, in both Ashkenazi and Sephardic samples, as well as in Samaritans and other religious minorities of the area, and the occurrence and absence of the relevant modal haplotypes, the CMH's, the GMH, the Bedouin Modal Haplotype, happen to fit the normative history of the region. Your treatment of the modern Palestinian Arab population as a millennia-old Palestinian genetic constant seems to be an assumption or axiom on your part, an axiom that doesn't require positive support, but only a series of sometimes dismissive ("it's logical to understand…", "…being a red herring") negative arguments.
maju:
"The true CMH"
I don't know if you're a biblical literalist, but if so, we're probably going to be talking past each other. The secular and historically-minded understanding is that the political situation in Judea of antiquity is that there have been a number of priestly families which have at differing periods held power and which we have no realistic reason to think all descend from a single person. The most recent paper on the CMH have found several Cohen Modal Haplotypes represented in different haplogroups. Thus we have several priestly lineages originating in a common time period and a common region such that these lineages would be represented in both Ashkenazi and Sephardic samples (that is, Judea, predating the formation of major diasporas).
"the J2 one being a red herring"
A red herring is a logical fallacy. The J2 CMH is a modal haplotype. I'm not sure what your purposes are in this discussion such that you would readily confuse the two.
"Canaanite is of course a relevant proto-historical term but they also had their genesis, which must have been rooted, Semitic invasions apart, on the Neolithic populations of the area."
Thus I take it you agree that the significant proportion of J2 existed in the Levant long before the Greco-Roman periods, and before the Jewish ethnogenesis. If the "early European admixture" of J2 has its origins in the Neolithic, then I have no problem with this in that it is consistent with the series of facts listed in my prior posting.
"Settlement? Can you refer to such documents?"
I'm referring to normative history. Alternative historical theses are interesting, but go outside the scope of this discussion. Geneticists are of course entitled and wise to interpret the genetic data under the normative understanding of historical events. Fred Donner's text discusses Arab-Muslim settlement in the early Islamic expansion, focusing mainly on Mesopotamia and to a lesser extent Syria/Palestine. Moshe Gil and others can be referred to for a more focused discussion of Palestine. Ramle of course was specifically an Arab Muslim settlement, and I'll mention the Arabian tribes of Bani Murrah, Bani Harith, Bani Salim, Bani Malik, Bani Hasan, Bani Judham, Bani Khawlan, Bani Ghassan, Bani Amila, Bani Khatham, Bani Himyar, Bani Kinda, Bani 'Amr, Bani Lakhm, Bani Thaqif, Bani Ghatafan, etc.
This isn't to suggest of course that sole settlement occurred during the initial Islamic expansion. Throughout the Islamic period there have been different periods of settlement. For example, Nimr's Tarikh Jabal Nablus discusses the wave of Turkic, Kurdish and Bedouin settlement in the Nablus, Jerusalem and Hebron areas since the 17th century. We learn from the various "chroniclers" of the 16th, 17th centuries and others (Volney, Burckhardt) that the area of Hebron was largely populated by descendants of bedouin tribes who immigrated there.
To mr "Kidproquo".
Jewish is a religion not an ethny.
The majority of Jews are not J2 nor have Cohen modal type.
You can not equate J2 with Jew or Hebrew.
Also I think that Arabs and other Arabianic peoples too should have this Cohen modal as they belong to the same Ethnolinguistic branch and Adnani Arabs are said to descend from Ismael son of Abraham.
Not all Palestinian Arabs are tribalized Arabs, and those ones are most likely the closest folk to the pre "jewish", and "jewish" folk of Israel/Palestina.
Israel can not stand isolated and animous toward their neighbors, it's only a matter of time; the early you become a "normal state" the better is for all.(remember South Africa)
But I think you know it's surrealistic and absurd to hang on some proteins (ie this Cohem modal haplotype) present in rather small portion of Jews.
With that logic, all "non native americans" etc... should go back to the old world.
We have a short live, let's live together in peace, brotherhood and harmony, whatever our race, language, religion or "modal haplotype" our diversities is our richness.
Finally as the great poet Nazım Hikmet said.
"güzel günler göreceğiz çocuklar"
Best regards for a peaceful and united world!
Maju - you raise a very good point that the previous intelligibility does provide a proxy for linguistic shift.
Also correction on Anatolia, but the Arabs did conquer the Kurds and the Caucasus.
I guess there are no geographic barriers between Arabia and North Africa - sort of morphed into one continuous political unit, whereas Iran, Caucasus & Anatolian are the highlands.
The only arabic speaking province in Iran is contiguous to south Iraq and are low-level desert plains (which incidentally are where the oil of Iran is primarily located).
From Wikipedia:
"The authors speculated that when the Assyrians conquered the northern kingdom of Israel, resulting in the exile of many of the Israelites, a subgroup of the Israelites that remained in the Land of Israel "married Assyrian and female exiles relocated from other conquered lands, which was a typical Assyrian policy to obliterate national identities."[26] The study goes on to say that "Such a scenario could explain why Samaritan Y chromosome lineages cluster tightly with Jewish Y lineages, while their mitochondrial lineages are closest to Iraqi Jewish and Palestinian mtDNA sequences." Non-Jewish Iraqis were not sampled in this study, however, mitochondrial lineages of Jewish communities tend to correlate with their non-Jewish host populations, unlike paternal lineages which almost always correspond to Israelite lineages."
I think trying to infer historical conclusions from genetics, particularly when at times the reams of data contradict each other, just doesn't work, sort of like grasping at straws.
It is well-known that the Arabs founded urban centres; the spread of Islam coincided with the spread of cities and urbanisation in Iran.
However one must really beg the question how many Bedouins were there and how were they able to expand so much faster than the natives.
Has anyonetested the Palestinian Christian population; they would be the best test case since they would have no Arab-Islamic influx?
How about being specific to the report and not resorting to mumbo jumbo and religious concepts. I doubt anything written or said about the ancestors of the people we call Jews. In other words, no 12 twelves, no priests, no Aaron, no Moses - it is all just religious clap trap.
Now with regards to this study. I want to know what the colors represent in the last frappe. Dienekes said the purple does not mean it is Mozabite Berber because it is highest in the Mozabites but it is found in other groups like the East Asians. Well what is it?
Re. CMH. Remember that there was a genocide in Roman times and that while Christians were spared, Jews in general and, with all likelihood, specially their elites, including Cohanim, were targeted.
"Under what rule of logic do you make the inference that Cohanim remained within the Jewish faith but non-Cohanim didn't?"
Cohanim were (are?) the priestly caste, their whole lifestyle depended on religious adscription. Not surely comparable but you may correlate with modern conversions of Hindus to other religions, which are almost exclusive of the low castes and not any Brahmin phenomenon.
It seems only logical. And even more if we consider the speculated relations of early Christians (mostly Jews) with dissident sects within Judaism such as the Essenians, which challenged the religious and social monopoly of the Malachite priestly caste.
It's almost impossible to reconstruct what exactly happened then but what I'm saying here has some good logic at least. Remember that Christianism was at its origins a Jewish sect and that it needed of several centuries to become powerful enough to take over the Roman Empire. Time in which it was surely difficult to differentiate between Christians and Rabbinic (or whatever else) Jews, as all them were proselytist and had strong presence in West Asia, specially in Asia Minor. Only when Christians took over the Empire we can really begin to see them as clearly different (mainstream) from other Jews (minority).
"Not to invoke an anthropological platitude, but nomads are more likely to eventually become settled populations than settled populations are likely to become nomads. And more so, Arab Muslim bedouin are more likely to become settled Arab Muslims than Rabbinical Jews are likely to become Arab Muslim bedouin".
Can't but agree with this (laughs). But count the nomad Bedouins and the settled Muslims in modern Palestine/Israel. Bedouins are necessarily a small minority always for reason of their lifestyle and therefore they can't substantially alter the sedentary genetic landscape even if all would decide to settle at once.
"the significant proportion of J2, in both Ashkenazi and Sephardic samples, as well as in Samaritans"...
Modern Samaritans are a tiny population of 712 individuals (2007 data) and historically they were immigrants from possibly Iraq (or in any case further north/NE where J2 is much more abundant).
"the CMH's, the GMH, the Bedouin Modal Haplotype"...
They are probably just common local haplotypes. The CMH is not even something exclusive of Cohanim nor Jews, not at all, and most Cohanim have other haplotypes. Somewhat speculative at least.
"I don't know if you're a biblical literalist"...
I'm not (please!), I just give the benefit of doubt to legends when they seem somewhat plausible.
"A red herring is a logical fallacy. The J2 CMH is a modal haplotype".
I'm just supposing that there is a real Aaron-founded lineage, with the other lineages being co-opted or illegitimate children in Cohen families (est. 1/10 children is, most never get to know). But you may well be right that the legend has no real foundations. I can concede on that.
"Thus I take it you agree that the significant proportion of J2 existed in the Levant long before the Greco-Roman periods, and before the Jewish ethnogenesis".
In the Levant yes but I would say that not really so much in Palestine but rather in Syria-Lebanon. J2 has a clear more northernly ("highlander") distribution than J1 and is precisely one of the reasons why I suspect an Anatolian origin of at least part of the modern Western Jewish ancestry.
The hypothesis that there was a population replacement in Palestine (of all places) does not stand on light of the North African connection of J1, which is obviously very old and must be original from West Asia in or around Palestine. You can't really claim an Arab origin of North African J1 and there's nearly no J2 in North Africa either. This distribution pattern of the two J sublineages must have very old roots. I think it must be from Paleolithic times but even if it is Neolithic it would not make any difference for this debate. Palestine is the nexus between North African and West Asian J1, which must have gone forth and back between the two regions on light of haplotype structure (but ultimately has West Asian origin).
"I'm referring to normative history".
Sorry, I don't know what "normative history" means and could not find it in Wikipedia.
"Fred Donner's text discusses Arab-Muslim settlement in the early Islamic expansion"...
I won't question that. What I say is count the bedouins and count the sedentary residents in any of these countries. It is blatantly obvious that the genetic impact of any settlement of nomads must have been very limited.
You cannot make up a purely or almost purely Bedouin ancestry of Palestinians out of such a limited settlement, exactly the same you can't pretend that Spaniards descend from the Visigoths or Tunisians from the Vandals (except to a very minor extent).
It is simply unbelievable. If there was ever a (necessarily partial) population replacement of some entity in Palestine it must have happened in Roman times after the Jewish wars. We have no documents that tell us of that but I could concede that it's not totally impossible considering the extent of the Jewish genocide in that time. However Palestinian genetics don't seem to support it.
"Your treatment of the modern Palestinian Arab population as a millennia-old Palestinian genetic constant seems to be an assumption or axiom on your part, an axiom that doesn't require positive support"...
There is positive support in historical data and in all the genetic data I have been able to see (though Palestinians are comparatively under-researched). I can't say if there was or not some sort of demic replacement in Roman era but for sure that there was not such thing with the Arab/Muslim invasion. That's just a Zionist neo-myth, part of the hasbara to justify the invasion and robbery of the Palestinian land.
Ironically enough, early Zionists ideologues knew very well that Palestinians had to be at least to a large extent descendants from historical Jews. And I agree with them in this: peasants stick to their land much more than they stick to their faith or language.
Even when they emigrate, they also remain. Irish may have emigrated en masse to half the world but they are still the vast majority of people in Ireland too. Is this an axiom? It is just the null hypothesis and so far it does not seem that anything contradicts it.
Actually this paper shows that Palestinians are closer (Fst) to Western Jews than any other considered population but (oddly enough) North Italians and almost as close to them as Western Jews are to each other on average. And are certainly closer than Iranian and Iraqi Jews are. In fact, I think that the labels in the graph posted here are inverted for Palestinians and Bedouins, because Bedouins are much more distant Fst-wise from Western Jews (or Jews in general, or any other considered population too) than Palestinians are:
West Jews with each other (average): 0.005
West Jews - Palestinians: 0.008
Palestinians - Bedouins: 0.009
West Jews - Iraqi Jews: 0.010
Palestinians - Iraqi Jews: 0.010
West Jews - Iranian Jews: 0.016
Palestinians - Iranian Jews: 0.017
For comparison, the most distant European samples (Basques and Russians) have Fst=0.016 in regard to West Jews (averaged) and Fst=0.021 in regard to Palestinians.
ashraf:
"The majority of Jews are not J2 nor have Cohen modal type."
When did I claim such? I hope you're not trying to strawman me here. The point of the discussion of the Cohen Modal Haplotypes is to point out that 1) we have genetic data that supports the historical data regarding a common regional pre-diaspora origin of the major diaspora Jewish groups, and 2) the contention that the genetic landscape of Palestine was not genetically altered during the Islamic period is apparently untrue (and this happens to also fit the historical data).
"You can not equate J2 with Jew or Hebrew."
When did I do such? I pointed out that the significant presence of J2 in the Levant probably predated the Greco-Roman period.
"Also I think that Arabs and other Arabianic peoples too should have this Cohen modal"
I don't know what an "Arabianic" person is supposed to be, but as it turns out, the extended Cohen Modal Haplotypes is almost entirely restricted to Jewish populations. Therein lies its relevance to the theses described below
"they belong to the same Ethnolinguistic branch and Adnani Arabs are said to descend from Ismael son of Abraham. "
I take it that like maju, you are a biblical (or koranic?) literalist?
"Not all Palestinian Arabs are tribalized Arabs"
I didn't assert such. In fact I've been at pains to point out the multiple origins of the Palestinian Arabs that have taken place throughout the Islamic period. I can name several PA clans of Christian or Jewish heritage. Maju has asserted explicitly or implicitly three theses:
1 Muslims did not genetically alter the landscape of the middle east
2 Contemporary Paletinian Arab population represents a kind of ancient control sample against which everything else is tested against for, say, 'levantiness' (or 'palestiness'?)
3 groups not sufficiently similar to this control sample have no origin in the levant (although he applies this in contradictory ways--Druze are "immigrants," Maronites are not, Mizrahim are, Samaritans are who knows what?)
All of these theses are apparently false in light of both the genetic and historical data. Only major qualifications will salvage anything.
The rest of your political claptrap is ironic. I've only been defending the normative historical view, and the geneticists' own interpretations of their own work (!) It appears that maju is at pains to present substantially alternative interpretations, which he is entitled to do, so direct your ascription of political motive his way.
Mr AAron and Zachary.
Of course there was language replacement but the "Arabic" influx was not insignifiant(compared to let's say Angola which is now majoritly Lusitophone but with very few Portuguese influx).
If you look very well you will see that All countries with more than 20% J1-YDNA are majoritly (ie more than half)Arabic speaking (except perhaps Israel, but Israel is majoritly Arabianic speaking anyway[ie Hebrew+Arabic]and was majoritly Arabic speaking 70 years ago).
I think that early Arabs "did not want" that the other non Arab folks become Arabic speaking and Muslims(a totally different policy than the hegemonious romanisation of western Europe both culturally and linguistically).
Also dont forget that Sumerians and Elamites assimilated into Akkadians and Persians well before the rise of Islam.
"The hypothesis that there was a population replacement in Palestine (of all places) does not stand on light of the North African connection of J1, which is obviously very old and must be original from West Asia in or around Palestine. You can't really claim an Arab origin of North African J1 and there's nearly no J2 in North Africa either. This distribution pattern of the two J sublineages must have very old roots. I think it must be from Paleolithic times but even if it is Neolithic it would not make any difference for this debate. Palestine is the nexus between North African and West Asian J1, which must have gone forth and back between the two regions on light of haplotype structure (but ultimately has West Asian origin)."
In North Africa,It spread to North Africa (as identified by the motif YCAIIa22-YCAIIb22; among Algerians 35.0%, Tunisians 31%), J1 first entered Ethiopia with the spread of Semitic speakers [citation needed] Eritrea (11%), Ethiopia (9%), Ethiopia-Amhara (33.3%). J1 also may be found with high frequency in the northern parts of Sudan (J-12f2(xJ2-M172): Arabs 45%, Nubians 41%, Copts 39%, Beja 36%), and present with lower frequency in the region of Darfur (J-12f2(xJ2-M172): Masalit 6%, Fur 6%).[17] Haplogroup J1 may be found in as many as 20% of Egyptian males,[18] with the frequency of this haplogroup tending to be comparatively high in the south of the country.[19]
good point Maju when in doubt use the equivalent Christian populations as a control; Egyptian Copts are most definitely NOT the descendants of Bedouins (at least as a result of the Arab-Islamic invasions).
The Christian Arab populations (of which Syria, Lebanon, Palestine, Egypt, Iraq all have in abundance @ 5-10%; Lebanon obviously higher) are great ways to test for "Arab-Islamic" demographic expansion.
In the Muslim Maghreb the Berbers are a great control to test the extent of ancestry.
For the Khaleej; Gulf Arabs, I think its pretty well substantiated that's its a different ancestral pool (Bedouin etc.)
This seems to be a pretty nifty tool; two criticism doesn't drill into the haplotypes and the other "national" populations don't make much sense in arbitrarily defined states.
http://en.wikipedia.org/wiki/Y-DNA_haplogroups_by_ethnic_groups
@Zachary: the main reference in this regard is surely Semino 2004. Table 2 provides frequencies that support and N/S distinction in West Asia for J2/J1 and also shows that, in concordance with this division, J1 is rare in Europe and J2 is rare in Africa, suggestive of an even sharper divide at some time in the past.
But most important for me is the haplotype structure in figure 4, that shows how intermingled is the bulk of J1 (J1* but it's almost all J1 in this paper), in particular the shaded area, characterized by the YCAIIa-22/YCAIIb-22 motif, and which includes most individuals within the haplogroup.
This suggests that most J1 spread in a process that heavily involved North Africa and that might even have originated there, rather than West Asia, possibly within the proto-Neolithic Afroasiatic (?) expansion, that also carried E1b1b1 to West Asia and Europe.
Back to frequencies, we can see that percentages as high as Jewish of J2 are only found North and NE of Palestine (Lebanese, Iraqi, Kurds, Konya Turks, Georgians), as well as in SE Europe (specially North Italians). By subhaplogroups however, the most similar may well be Konya Turks but these cannot account for the high apportion of Jewish J1 which may well be of genuine Palestinian (ancient Jewish) origin. Not written on stone anyhow, just my best guess.
maju:
"Re. CMH. Remember that there was a genocide in Roman times and that while Christians were spared, Jews in general and, with all likelihood, specially their elites, including Cohanim, were targeted."
A genocide of Palestinian Jews in Roman times would mean that, contrary to your implication in your prior posts, contemporary Palestinian Arabs are not "the most genuine descendants of ancient Jews." The inconsistency needn't trouble us since the genocide was specific to Jerusalem not Palestine as a whole. Jews, at least together with Samaritans, constituted the majority in Palestine at the eve of the Islamic expansion (see Gil).
"It's almost impossible to reconstruct what exactly happened then but what I'm saying here has some good logic at least."
Unfortunately, without ushering in authorities, what you're doing is engaging in speculation. Your contention that Cohanim are immune to the same historical forces to which non-Cohan Jews are subject is unusually ad hoc.
"Modern Samaritans are a tiny population of 712 individuals (2007 data) and historically they were immigrants from possibly Iraq"
It is interesting that you allow for Samaritan immigration into Palestine but not for Arabian (or other Muslim) immigration into Palestine. One could invoke all of the exact same precepts that you assume and which form the basis of your contention that PA samples are unexceptionally representative of ancient the Palestinian population to similarly assert that Samaritan samples are unexceptionally representative of the ancient Palestinian population. A more rigorous attempt at consistency on your part would be helpful. The claim that Samaritans are immigrants from Iraq comes from the same historical sources which assert that Samaritans are the patrilineal descendants of the indigenous population of Samaria. Your endorsement of one half and rejection of the other half of the same narrative is again ad hoc and inconsistent on your part.
I will give qualified credence to both relevant narratives in interpreting the genetic data: that Samaritans have their origins in Samaria, and Palestinian Arabs have origins in Arabia (according to traditional Muslim-Arab historiography).
"The CMH is not even something exclusive of Cohanim nor Jews"
The original CMH occurs as a singleton in Palestinian Arab samples, and in moderate frequencies in some other populations. The extended CMH occurs in PA and other non-Jewish samples not at all. A problem for the claim that Muslim settlement into Palestine did not alter the genetic landscape of Palestine.
You seem to be ascribing to me the view that there has been some sudden massive population displacement in the 7th century in Palestine. I have not asserted this, but only pointed out what exists in traditional Muslim historiography that Muslim settlement in Palestine has occurred throughout the Islamic period.
I had to google "hasbarist" since it's a retort you've used twice so far. You're entitled to believe that early Islamic Arabic authors (e.g. Tabari), 17th century chroniclers, and modern Arabic-language historians (e.g. Nimr, Shihabi) were employed by the state of Israel, but I have to express skepticism ;)
"I'm just supposing that there is a real Aaron-founded lineage, with the other lineages being co-opted or illegitimate children in Cohen families"
Even the Bible and contemporary biblical criticism asserts the existence of unrelated powerful priestly families. My point was merely that the incidence of the multiple modal haplotypes associated with a ancient Jewish tradition and found in major diaspora samples supports the common regional pre-diaspora origin of the relevant populations, and refutes the claim that Muslim settlement in Palestine did not alter the genetic landscape.
"In the Levant yes but I would say that not really so much in Palestine but rather in Syria-Lebanon."
And your drawing of an invisible magical border just south of Tyre that bars further diffusion of J2 in the neolithic is just as ad hoc as your prior claims. There is more than enough archaeological and epigraphic evidence of the Canaanite presence in Palestine.
"This distribution pattern of the two J sublineages must have very old roots. I think it must be from Paleolithic times but even if it is Neolithic it would not make any difference for this debate. Palestine is the nexus between North African and West Asian J1, which must have gone forth and back between the two regions on light of haplotype structure (but ultimately has West Asian origin)."
I'm not sure what the point of your discussion here is. Of course J1 was substantially represented in ancient Palestine. The modal haplogroups represented in ancient Palestine would of course include J1, J2, and some subclades of E, with a few other smaller elements. And it is interesting that the genetic data supports that this distribution is represented in the Samaritan population almost perfectly.
"Sorry, I don't know what "normative history" means"
As in, not revisionist history. Though revisionist history typically comes from revisionist historians, and you've yet to invoke historians of any sort at all.
"You cannot make up a purely or almost purely Bedouin ancestry of Palestinians"
My examples of Muslim settlement in Palestine at different periods throughout the course of 13 centuries of the Islamic period in a prior post have done nothing of the sort.
Lebanon (Capelli et al. 2005; comprised of 43 Christians and 39 Muslims)
14/82 = 17.1% E1b1b
Ashkenazim Jewish (Semino et al. 2004)
1/77 = 1.3% E1b1b1-M35(xE1b1b1a-M78, E1b1b1b-M81, E1b1b1c-M123)
4/77 = 5.2% E1b1b1a-M78
9/77 = 11.7% E1b1b1c-M123
14/77 = 18.2% E1b1b total
Lebanon (Semino et al. 2004)
5/42 = 11.9% E1b1b1a-M78
1/42 = 2.4% E1b1b1b-M81
2/42 = 4.8% E1b1b1c-M123
8/42 = 19.0% E1b1b total
Ashkenazi Jews (Behar et al. 2004)
71/442 = 16.1% E1b1b1-M35(xE1b1b1a-M78, E1b1b1b-M81)
12/442 = 2.7% E1b1b1a-M78
4/442 = 0.9% E1b1b1b-M81
87/442 = 19.7% E1b1b total
Ashkenazi Jews of Polish origin (Shen et al. 2004)
2/20 = 10.0% E1b1b1c1-M34(xE1b1b1c1b-M290)
2/20 = 10.0% E1b1b1a-M78(xE1b1b1a1a-M224)
4/20 = 20.0% E1b1b total
Arabs from Tanta and Cairo, Egypt (Luis et al. 2004)
4/147 = 2.7% E1b1b1-M35(xE1b1b1a-M78, E1b1b1b-M81, E1b1b1c-M123)
26/147 = 17.7% E1b1b1a-M78
12/147 = 8.2% E1b1b1b-M81
10/147 = 6.8% E1b1b1c-M123
52/147 = 35.4% E1b1b total (Luis et al. 2004)
"But, the basal E1b1b1*/M35* subclade is found in 20% of the Ashkenazi Jewish population, suggesting that it is a part of the founding population of Jews."
E1b1b1-M35(xE1b1b1a-M78, E1b1b1b-M81, E1b1b1c-M123) has been found in Ashkenazi Jews in commercial testing in addition to one individual in the Ashkenazi sample of Semino et al. 2004, but it is not that frequent among them. The majority of E-M35 Ashkenazim belong to the subclade E1b1b1c-M123.
I said,
"Haplogroup E3b generally is rather uncommon in populations of Arabia, so I think the neutral expectation is that E3b should not occur with particularly high frequency among Bedouins. E3b actually is significantly more common among settled populations of the Nile Valley than it is among recently nomadic or semi-nomadic populations of the Arabian Peninsula."
In response to my previous comment, Andrew Oh-Willeke said,
"E3b and Mozabite contributions generally are common to the same extent among the Levantine populations studied in the linked paper (Jews, Druze, Palestinians). There are more Mozabite contributions are more common in Bedouins than they are in Levatine Arab/Druze populations suggesting a longer period of admixture."
Claiming that a greater percentage of the males of an ethnic group must belong to a particular subclade of a Y-DNA haplogroup because a sample of individuals from that ethnic group has exhibited greater membership in a particular cluster produced in a STRUCTURE analysis of autosomal data is utterly ridiculous.
Andrew Oh-Willeke said,
"E3b is present at significant frequencies in Arabia in two of its subtypes."
Not really. On average, E3b probably is no more common in Arabia than it is in Europe and West Asia outside the Levant. Levantine populations (including Jews) tend to have somewhat higher frequencies of E3b, with the Bedouins being perhaps intermediate between Arabians and Levantines in this respect. However, Egyptians have even more E3b than Levantines. On the other hand, Sudanese Arabs, who generally are much more nomadic (or at least recently descended from nomads) than Egyptian Arabs, seem to have oddly low frequencies of E3b:
Qatar (Cadenas et al. 2008)
1/72 = 1.4% E1b1b1a2-V13
1/72 = 1.4% E1b1b1a3-V22
1/72 = 1.4% E1b1b1a-M78(xE1b1b1a2-V13, E1b1b1a3-V22)
1/72 = 1.4% E1b1b1c1-M34
4/72 = 5.6% E1b1b total
Saudi Arabia (Abu-Amero et al. 2009)
4/157 = 2.5% E1b1b1-M35(xM78, M81, M123)
1/157 = 0.6% E1b1b1a3-V22
7/157 = 4.5% E1b1b1c1-M34
12/157 = 7.6% E1b1b total
United Arab Emirates (Cadenas et al. 2008)
1/164 = 0.6% E1b1b1a2-V13
11/164 = 6.7% E1b1b1a3-V22
1/164 = 0.6% E1b1b1a-M78(xE1b1b1a2-V13, E1b1b1a3-V22)
1/164 = 0.6% E1b1b1b-M81
5/164 = 3.0% E1b1b1c1-M34
19/164 = 11.6% E1b1b total
Yemen (Cadenas et al. 2008)
1/62 = 1.6% E1b1b-M215(xE1b1b1-M35)
2/62 = 3.2% E1b1b1-M35(xE1b1b1a-M78, E1b1b1b-M81, E1b1b1c-M123)
5/62 = 8.1% E1b1b1c1-M34
8/62 = 12.9% E1b1b total
Omani Arabs (Luis et al. 2004)
2/121 = 1.7% E1b1b1a-M78
15/121 = 12.4% E1b1b1c-M123
17/121 = 14.0% E1b1b total
Bedouins (Cruciani et al. 2004)
1/28 = 3.6% E1b1b1a-M78 (Delta cluster)
1/28 = 3.6% E1b1b1b-M81
2/28 = 7.1% E1b1b1c1-M34
4/28 = 14.3% E1b1b total
Lebanon (Zalloua et al. 2008)
2/916 = 0.22% E1b1b1-M35(xE1b1b1a-M78, E1b1b1b-M81, E1b1b1c-M123, E1b1b1e-V6) [Muslim]
96/916 = 10.48% E1b1b1a-M78 [most frequent among Druzes]
11/916 = 1.20% E1b1b1b-M81 [most frequent among Druzes]
39/916 = 4.26% E1b1b1c-M123 [most frequent among Christians and Muslims]
148/916 = 16.2% E1b1b total [seems to occur most frequently among Druzes]
Sudanese Arabs (Hassan et al. 2008)
6/102 = 5.9% E1b1b1a1-V12(xE1b1b1a1b-V32)
7/102 = 6.9% E1b1b1a1b-V32
4/102 = 3.9% E1b1b1a3-V22
17/102 = 16.7% E1b1b total (all derived from E1b1b1a-M78)
Kidproquo: I think that the new paper by Behar pretty much answers our discussion. Western Jews in particular cluster best (almost identical in the K-means analysis) with Lebanese and Cypriots, while Palestinians appear at K=8 as a clearly different population to a very large extent from Bedouins, Arabs, modern Jews of all kinds and any other population studied.
This seems to address the issue of the possible demic replacement in Palestine in a negative sense, because no source population is apparent at all. Anyhow, for what I know of the Roman genocide it may not have been so extremely systematic after all and certainly Jewish Christians were spared.
"I had to google "hasbarist""...
Hasbara: it means propaganda in Hebrew. Admittedly I did not know this word a couple of years ago either.
"And your drawing of an invisible magical border just south of Tyre that bars further diffusion of J2 in the neolithic is just as ad hoc as your prior claims".
It's not me who is drawing the border but all the available genetic data. It's not a strict division (i.e. lineages have obviously migrated between the two subregions afterwards) but it is certainly a quite clear one. My point is that Palestinian J1 is not (mostly) of Peninsular Arabic origin. The opposite instead is very possible.
"It is interesting that you allow for Samaritan immigration into Palestine but not for Arabian (or other Muslim) immigration into Palestine".
I don't deny Arabian or other Muslim immigration into Palestine, what I say is that it was not so important as to change the genetic landscape in the manner you wishfully think. And the new data of Behar clearly ratifies this fact: a sizable fraction of the Palestinian sample show only the specific Palestinian component at K=8, while the rest do show variable admixture with other Arabs, which may be of Muslim or more ancient periods (the "Arab" component is widespread, including among Jews and, in smaller amounts, some Europeans, meaning it doesn't need to be strictly Arab but just the main West Asian lowlands component).
Samaritans are also high in this "Arab" component.
The Palestinian pattern showing an specific ethnic-genetic component totally dominant among many of them but admixed among others is similar to that of Druzes and Negev Bedouins, however each of these groups has their own unique ethnic components, meaning that they are unique distinct populations.
This relative inbreeding of Palestinians is highly contradictory with your claims that that they are nothing but undifferentiated Arab immigrants. The data of Behar does not support your claim at all but clearly says that there is distinct genetic component specific of sedentary Palestinians that has been variedly diluted or, in many cases, not diluted at all.
Live to learn...
@ Zachary Latif
"is there a haplotype specifically associated with the Arab-Islamic conquests of the 7th century."
If there is, it is quite thin. The Islamic empire expanded from greater Mecca to include an area including Syria, Mesopotamia, Egypt and Lybia from 636 CE-646 CE, and conquered Persian from 637 CE to 639 CE. Mass conversion was the rule and the bureacratic classes of rome and Persia were mostly left in place. The lion's share of the force that conquered Visigothic Spain in 711 CE was made up of Berber converts from the general vicinity of Algeria and Morroco. But, by 1000 CE, the Islamic empire had fragmented into a mostly religiously united but ethnically divided confederation of states rather akin to the Holy Roman Empire after Charlamagne's death. While many of the Muslims would made up the bulk of the demic force of Islamic expansion in North Africa and Iberia were ethnically Berber, many of the Muslims involved in the Islamic expansion in the East were ethnically Turkish.
South Asia shows little sign of any ethnic contribution of invaders after the Indo-Aryans. Neither the Greek nor the Muslim invaders had nearly as significant a genetic impact.
The Islamic expansion, genetically, looks more like the Hungarian language speaking people's invasion that created Hungary, but left little genetic trace in the conquered peoples.
maju:
"... is highly contradictory with your claims that that they are nothing but undifferentiated Arab immigrants"
I don't mean to be rude but I'm flabbergasted that this the third time you've strawmanned me with that. I've specifically referenced multiple origins of immigration into Palestine in terms of time periods, geographic regions, ethnic or population types. I've referenced the significant Kurdish and Turkic (Circassian, Turkoman, Mameluk) immigration into Palestine (groups that themselves became "Arabized") during the post-Crusader period. Large PA clans today are traceable to such immigration...the Tuqans, Nashashibis, Jayyusis, Doghmushes, Abu Goshes, etc. And your use of "Arab" in such an undifferentiated way betrays your ahistorical assumptions. You ignore the historical realities that would give rise to the relative genetic distinctiveness amongst various "Arabic" groups, and the history of their immigration into Palestine. Bedouin versus sedentary populations. Hijazi versus Nejdi versus Syro-transjordanian versus South Arabian/Yemeni (Yemenis themselves being historically a culturally "Arabized" people). These distinctions exist in the tribes I've previously listed.
"It's not me who is drawing the border but all the available genetic data."
Something you've repeatedly asserted rather than demonstrated.
"Hasbara: it means propaganda in Hebrew."
So…Tabari, Volney, Buckhardt, Nimr, etc, are Israeli propagandists? Or they aren't? This is becoming silly.
"I don't deny Arabian or other Muslim immigration into Palestine, what I say is that it was not so important as to change the genetic landscape in the manner you wishfully think."
You've stated that Muslim immigration into Palestine could not alter the genetic landscape in any meaningful manner. I don't know what a meaningless genetic alteration would be beyond your previous prior allusion some non-settling conquerors and missionaries. That is a fairly explicit denial of Muslim and Arabic immigration into Palestine, a wishful and unsupported revisionism on your part that contradicts the historical data.
I don't have the recent Behar paper so I can't comment on it. I don't know what you're referring to as the "Arab component" but I doubt this is a descriptor the authors have endorsed. A descriptor that researchers have endorsed however is the low-diversity "Arab clade," which is substantially represented in PA samples, and demonstrably of non-Palestinian origins. Another example of Arab-Muslim meaningful genetic alteration.
@Kidproquo:
"I've referenced the significant Kurdish and Turkic (Circassian, Turkoman, Mameluk) immigration into Palestine"
Those immigrations are not apparent at all in Behar 2010 Palestinian autosomal data. Circassians anyhow are still a distinct population in Palestine today and they have sworn loyalty to Israel (they are the only minority serving in the Israeli Army). These documented migrations do not seem to have meant any significant impact on Palestinian genetics.
I insist that you take a look at Behar's paper (discussed here at Dienekes) and specially at the K-means analysis of all Jews and most West Eurasians. It's obvious that Palestinians are pretty much unique in this subcontinental context and not the product of any recent immigration.
And this really settles the debate for good.
"And your use of "Arab" in such an undifferentiated way betrays your ahistorical assumptions".
The use of the term "undifferentiated Arabs" Reflects your pseudo-historical claims and was meant to indicate your point, not mine:
"... your claims that that they are nothing but undifferentiated Arab immigrants".
I'm the one to be "flabbergasted" at your confusionist manipulation of my words, if anyone.
"So…Tabari, Volney, Buckhardt, Nimr, etc, are Israeli propagandists?"
No idea, I haven't read them and mostly they are unheard names for me. I doubt they are geneticists.
"... a wishful and unsupported revisionism on your part"...
I used to be pretty much into history and I have never before today, except from the mouth and pen of Israeli propagandists, read anything clearly suggestive of any clearly strong Islamic colonization in general much less of Palestine of all places.
But the discussion goes clearly beyond history, because it's obvious that in history often the anecdote is more relevant than the bulk of reality, that a 1% of aristocratic warriors have got many many more pages than the 90% of established peasants, whom most historians of the past and even today, don't seem interested to write about and, even when they do, don't seem able to find many materials that could tell something.
That is why population genetics is so valuable on its own right. And that is why I insist that you bother looking at the relevant genetic data.
"I don't have the recent Behar paper so I can't comment on it".
The relevant graphs are all available at the free supplementary material and have been posted by Dienekes (link above) and by myself.
Take a look and open your eyes.
The pink-colored "Arab" component (in the Euro-Med graph) is likely to be sliced into more specific local/regional components if the K-means analysis would have gone further, as would be the case with other widespread components. It is not in itself evidence of intense genetic identity (but at least some). But what is most important is the Palestinian component (coded in dark red or maroon), which is almost exclusive of Palestinians and in many of them virtually the only component at all (meaning that they relate much more strongly with other people with that component than with the rest).
This novel element, specially because of the wide array of compared populations and the relative shallowness of the K-means (K=8) where it shows up, clearly indicates that the Palestinian genetic pool is pretty much uniquely Palestinian and not any recent arrival from anywhere else.
@Andrew:
Mostly agree, just a minor correction:
"... rather akin to the Holy Roman Empire after Charlamagne's death".
Actually the German Kingdom and, hence the HRE (its projection into Italy and Burgundy), was a very solid state and probably the main European power until the death of Frederick II in the 13th century. Otherwise your comparison surely applies to a large extent.
"Circassians anyhow are still a distinct population in Palestine today"
Another willful misreading of what I wrote. I was specifically referring to Arabized clans of Circassian and other origins, not the non-Arabized Circassians who immigrated in the 19th century.
"No idea, I haven't read them and mostly they are unheard names for me. I doubt they are geneticists."
I'll take this as a tacit admission that you aren't interested in the normative historical understanding of the relevant regions and people. Unfortunately the genetic data cannot be interpreted in terms of how the populations are identified except in the light of historical data. Ethnonyms like "Arab," "Jew", "Druze," "Bedouin" all have historical presuppositions. Obviously you don't really believe historical data is irrelevant, but given the number of historical inaccuracies (and inconsistencies) expressed in your posts, it might as well be irrelevant to you.
Again, I don't have the paper and so I cannot evaluate your claims against the authors' own detailed analysis of the data. I don't see anything called an "Arab component" in the abstract, so I suspect this is a descriptor of your own contrivance reflecting your own ethnic presuppositions. Nothing in the abstract or graphs appears inconsistent with the findings of prior publications, which show a clustering of PA samples with Bedouin and Arabian samples, but with a distinctiveness consistent with 14 centuries of geographic-based bottlenecking as well as post-Crusader admixture from non-Arabian sources. Specifically it is consistent with the findings of a substantial low-diversity "Arab clade" indicative of recent founding events.
"I'll take this as a tacit admission that you aren't interested in the normative historical understanding of the relevant regions and people".
I am interested in history, however historical interpretations vary a lot. I don't know why you insist in adding the adjective "normative" to the word history. As fas as I know, there is no such thing.
Whatever the case if some punctual migration was recorded in some source, that does not necessarily mean at all that it was massive or implied demic replacement. I know of many many "migrations" in my area and most have not left any apparent genetic trace at all. Certainly not any large one: not Celts, not Phoenicians, not Romans, not Alans, Vandals and Swabians, not Goths, not Arabs and Berbers either...
They are recorded in history but their genetic input is minimal, if any at all.
"Ethnonyms like "Arab," "Jew", "Druze," "Bedouin" all have historical presuppositions".
Large ethnonyms like Arab are nothing but linguistic descriptions: if you speak Arab from childhood you are an Arab. The same is true for the ethnonym Chinese, Spanish (or Hispanic), etc. These nations have assimilated many other nations in whole.
Jewish and Druze are mostly religious descriptions, at least historically. However, as these communities have been mostly closed to external inputs (conversions) from at least 1000 years ago, they have eventually evolved in also ethnic communities, what can't be said of most other religions.
Bedouin is a subgroup of Arabs, just like Palestinians, etc. Their main difference is lifestyle as desert and semidesert pastoralists (and occasional raiders). They seem to have a genetic distinction, at least least some of the Negev ones.
"Again, I don't have the paper"...
I don't have it either (yet) but I can draw very clear conclusions only on the supplementary material, specially supplementary fig. 4b, which is the rally revealing stuff.
There's a pink or light purple component which shows up since K=7 and is most common among West Asian Arabs and Yemenite Jews. That's what I call the "Arab" component.
However at K=8, there is another distinct component (color coded brown or dark red) showing up as a markedly distinct Palestinian component. Apart of Palestinians only Negev Bedouins and Jordanians show it at some less important levels, with just anecdotal presence among other groups of the Northern Levant, including Western Jews.
This clearly demonstrates that at least the backbone of Palestinian genetics is not from anywhere else we can discern (and the sample is very very extense this time) and that this Palestinian genetic backbone is not really apparent among Jews except at very low levels.
For me it's the definitive nail on the coffin on other theories, such as the one you are defending.
Normative history: The core of history that is largely accepted amongst credentialed authorities in the relevant areas. Revisionist history: in this context, history that you've been literally making up as you go along.
Where in these admixture analyses do we find an age estimate for your PA component? In contrast, we are able to estimate the age of the Arab clade by reference to its low haplotype diversity. Herein lies the necessity of interpreting the genetic data in light of the historical data. Both are consistent with migrations beginning with the Lakhmids and Ghassanids and culminating in the Islamic expansion, and subsequent geographic-based bottlenecking. The deaths involved in the crusades would have also contributed to the bottlenecking.
@Kidproquo:
I have the impression that you revise "normative history" to fit your preferences. No big deal, as every other person does, but I'm 100% sure, that you don't have demographic statistics from the centuries you are talking about, just anecdotal reference to the arrival of this or that clan or exotic group.
It'd be like deriving the ancestry of Europeans by the data of the arrival of the Roma (Gypsy) wanderers or the the Vikings conquests!
I don't know enough about the history of the Levant but I know enough about the history of Europe, to be perfectly aware that there's simply no good statistics till the 19th century and no approximative data of much value before the very last of the Middle Ages.
So you are appealing to the opinion of some historians that I have never heard of in most cases (and I have read some many history books, believe me, including some about Muslim expansion and all that) and you call that "normative" in the hope to get your opinion backed by the opinion of some alleged "authority" and that way avoid facing the reality.
It's ok. No problem. Just don't expect me to believe in your particular selection of interpreters of almost non existent original sources because you say so. Not when the genetic data says otherwise.
"Where in these admixture analyses do we find an age estimate for your PA component?"
Nowhere. Do you really think that Romanized and Arabized Palestinians were isolated enough to develop such a distinctive component on their own right? Do you think that 12 million Palestinians (5 living in Palestine) are just like a bunch of ultra-isolated an hyper-endogamous Druzes?
I do not. So the Palestinian genetic peculiarity must be older. How older? Unsure; I'm inclined for Neolithic (more parsimonious) but it could also be from the Hebrew period. In any case it's blatantly obvious that they must be essentially the true descendants from the Biblical Jews.
But anyhow, I have already discussed above what happens with the much related Y-DNA J1 and J2 cline/divide.
And anyhow, it's so incredible that you guys are pushing against Occam's Razor of demographic continuity, clearly supported by all available genetic data, just because of favoritism for the myth that modern Jews descend from ancient Jews, disregarding the mounting evidence that Jewish genetics relates much better with Northern Levant peoples than with Palestinians, that I really find it ridiculous.
A North Levant origin was something I did not expect either, as I was almost persuaded about an Anatolian one, but I quickly accepted it as soon as the data became available. Why? Because all the genetic data we have on Jews, and it's more than for almost any other population anywhere, points to the North of West Asia, not Palestine. And that is also the case of the HISTORICAL data as well, because we know that modern Jews originated in the Diaspora, not specifically Palestine, and we know they were in that pretty much unspecific and highly proselytist.
It's all nothing but a legend. I feel like arguing with a Creationist...
Palestinians are a recent variable mix, as the unevenness of their ancestral proportions clearly indicates.
Maju's idiotic insistence that the Palestinian-centered cluster is of "Neolithic origin" can't explain it's unevenness in Palestinians, it can't explain why it barely registers among non-Arabs (a 10,000-year genetic component being largely confined to a 3-4ky ethnic group, right...), and in fact has nothing in its favor except Maju's irrational politics that want to see in modern Palestinian Arabs the original inhabitants of Palestine.
"Palestinians are a recent variable mix, as the unevenness of their ancestral proportions clearly indicates".
Palestinians do not only belong to the Palestinian cluster, I have agreed with that since the beginning. But I'm talking here of their specificity, of the Palestinian-specific cluster.
"Maju's idiotic insistence"...
No need to insult.
"... that the Palestinian-centered cluster is of "Neolithic origin" can't explain it's unevenness in Palestinians, it can't explain why it barely registers among non-Arabs".
Nor among Arabs other than Palestinians (Negev Bedouis, the only ones to show it at somewhat greater levels, are Palestinians too, even if a somewhat distinct subgroup - this presence is the least I would expect).
But you don't need to insist once and again in my preference for them having that distinct origin dating maybe to Neolithic. It's irrelevant. What matters is that it can't be recent, it can't be of the last 2000 years.
Doing that, instead of addressing the subject as I'm describing it, you are just trying to dismiss it by not considering the other possibility: that it formed later but, in any case, in some prolonged period(s) when South Levantines were clearly distinct culturally, for example in the ancient Hebrew era (roughly Iron Age).
En fin. I'm getting pretty much bored of this circular discussion.
What matters is that it can't be recent, it can't be of the last 2000 years.
Your opinion is worthless if it's not backed up by data or coherent arguments.
Your one "argument" self-destructs logically as I have already explained: if you can't see the Palestinian-centered cluster forming in the last 2,000 years due to outbreeding, then you have a much harder battle demonstrating that it formed in a longer period of outbreeding.
Until you manage to present a rational argument, I insist that your posts represent a case of "idiotic insistence".
Sigh!
Let's see Dienekes:
FIRST is I said above about the fact that the last two milennia have been unusually cosmopolitan and hence are the least likely ones to have produce a distinctive cluster among Palestinians of all peoples.
SECOND is the fact that we are comparing West Eurasians, and nearly them all (excellent sample), a population that we know has been diverging for the last 50-40,000 years or so. At such a shallow level (K=8) that European and most West Asian populations are rather amorphous, poorly defined as such populations, Palestinians show up as distinctive. Sure, sample size surely helped but there are other populations with larger samples who don't show up at all.
There is, of course, ascertainment uncertainty on what timeframe does that cluster represent but it just cannot have coalesced recently, when the population was surely more exposed to admixture with neighbours and even remote immigrants as the already mentioned Circassians, etc.
I think anyone can understand this reasoning and, if appropriate, discuss it coherently and not using the argument of the fool: insulting and disqualifying.
maju:
"So you are appealing to the opinion of some historians that I have never heard of in most cases (and I have read some many history books, believe me, including some about Muslim expansion and all that) and you call that "normative" in the hope to get your opinion backed by the opinion of some alleged "authority" and that way avoid facing the reality."
Yes, I back up my opinion with the opinion of relevant authorities, while you've back up your opinion with nothing. Like I said, you've been literally making up history as you go along in a number of startling propositions:
1. That literally no Arabian Muslim has settled in Palestine.
2. That there has been a genocide of the entirety of Palestinian, not just Jerusalemite, Jewry by the Romans (except for, as you put it, "Jewish Christians").
And I could go on.
onur:
I'm not going to spend much time on your post because I suspect you haven't been following the discussion. If someone were to assert, for example, the Roman Empire never existed, or that Charlemagne was a Buddhist, then your friend may very well endorse such opinions under the limitless subjectivity of the matter, and that's fine. However most working historians excluding your friend recognize that there is a core confluence of historical opinion built up over space and time.
maju again:
"Do you think that 12 million Palestinians (5 living in Palestine) are just like a bunch of ultra-isolated an hyper-endogamous Druzes? "
What's your point? Apparently you do think this. This is exactly what you've been pushing this entire time. You've stated that there has been no genetic alteration of the Palestinian landscape. Interesting how you apply this precept in inconsistent ways.
"And anyhow, it's so incredible that you guys are pushing against Occam's Razor of demographic continuity, clearly supported by all available genetic data"
No one is pushing what you've claimed. However you've been pushing for the proposition that there has been no genetic alteration of the Palestinian landscape (outside of "meaningless alteration").
"These haplotypes and their one-step microsatellite neighbors constitute a substantial portion of the total Palestinian (29%) and Bedouin (37.5%) Y chromosome pools and were not found in any of the non-Arab populations in the present study. The peripheral position of the modal haplotypes, with few links in the network (fig. 5), suggests that the Arab-specific chromosomes are a result of recent gene flow. Historical records describe tribal migrations from Arabia to the southern Levant in the Byzantine period, migrations that reached their climax with the Muslim conquest 633–640 A.D.; Patrich 1995). Indeed, Arab-specific haplotypes have been observed at significant frequencies in Muslim Arabs from Sena (56%) and the Hadramaut (16%) in the Yemen (Thomas et al. 2000). Thus, although Y chromosome data of Arabian populations are limited, it seems very likely that populations from the Arabian Peninsula were the source of these chromosomes." (Nebel 2001).
The substantial, low-diversity Arab clade puts to rest your bizarre picture of the PA population as a millennia-old genetically-unaltered genetic constant.
FIRST is I said above about the fact that the last two milennia have been unusually cosmopolitan and hence are the least likely ones to have produce a distinctive cluster among Palestinians of all peoples.
First of all, your contention that the last two millennia have been "unusually cosmopolitan" is unfounded.
Second, you have still failed to explain how a "distinctive cluster" would survive two thousand years (or even worse ten thousand) of gene flow.
SECOND is the fact that we are comparing West Eurasians, and nearly them all (excellent sample), a population that we know has been diverging for the last 50-40,000 years or so. At such a shallow level (K=8) that European and most West Asian populations are rather amorphous, poorly defined as such populations, Palestinians show up as distinctive. Sure, sample size surely helped but there are other populations with larger samples who don't show up at all.
First of all, your contention that West Eurasians have been diverging for 50 thousand years is (a) irrelevant, (b) unfounded, and (c) probably wrong, as most genes in West Eurasians have shallower genealogies than 50k and hence population divergence times are even later.
The populations that show up as distinctive are all non-European Muslims with high levels of consanguinity; there is your answer, and not Palestinian exceptionalism, whereby in all of Eurasia only in Palestine is there a distinctive remnant of the indigenous inhabitants.
In fact, you shoot yourself in the foot by postulating that the indigenous inhabitants' genetic legacy would be preserved in a region you label as "cosmopolitan".
I think anyone can understand this reasoning and, if appropriate, discuss it coherently and not using the argument of the fool: insulting and disqualifying.
Yep, anyone can understand that your "reasoning" is filled with hand-waiving, special pleading, and that it is motivated by your politics.
onur:
Impudent? haha :)
Actually it was a reductio ad absurdum rebutting your excessively silly response to, and misinterpretation of, my invocation of a normative understanding of history in a discussion which I suspect you haven't in fact been following.
Kidproquo:
"... a number of startling propositions:
1. That literally no Arabian Muslim has settled in Palestine.
2. That there has been a genocide of the entirety of Palestinian, not just Jerusalemite, Jewry by the Romans (except for, as you put it, "Jewish Christians")".
You have totally misunderstood what I said.
I can't but agree that some Arabian Muslims (and surely Arabized Muslims from elsewhere, like Syria, Egypt or Iraq) settled in Palestine. What I say is that there are no grounds and seems most unlikely that there was a massive settlement, much less a displacement of the original population in the Muslim period.
As for the Roman genocide, I haven't said that all Palestinian natives were exterminated. In fact now I'm pretty sure they were not. The only thing I said is that it was a much more clear opportunity, considering the dimensions reported by Roman age historians for this genocide (which of course can be exaggerated and distorted) for a demic replacement, if there ever was one.
As you insist that I mention my sources, F. Josephus, the main (only?) original source on this matter, talks of 1.1 million killed in Jerusalem alone. However it's very likely that the countryside was largely spared. This seems even more certain considering the new genetic evidence from Behar 2010 on modern Palestinians.
It would be interesting to study genetically Palestinian Jews (i.e. the descendants of those living in Palestine before the advent of Zionist immigration, which were some 10% of the population in the 1930s). Admittedly some of them may have arrived at different times in pilgrimages and such but they could still preserve some of the ancient Jewish/Palestinian genetics in them. I don't know if they keep some ethnic specificity in modern Israel but what I know is that no geneticist has paid them any attention so far.
I'll take a look to Thomas 2000 (if open access) but, from your quote, it seems to me that he's not talking of the bulk of their haplotype diversity but to a fraction of it (29%) which may indeed represent some historical expansion (not necessarily the Arab-Muslim one) with some founder effects. An Y-DNA expansion/founder effect affecting 29% of the Y-DNA pool may well mean just some 10% or even less of the whole gene pool, considering the peculiarities of gender-bias in demographic processes.
Look Dienekes, I'm pretty much tired of "discussing" with you in this matter because you are not discussing nor debating but mostly being nasty.
Still:
"... your contention that the last two millennia have been "unusually cosmopolitan" is unfounded".
Belonging to the Hellenistic states, the Roman Empire and then Islam makes this period pretty much cosmopolitan quite obviously. This is probably not a particularlirity of Palestine but also applies to that country.
"... you have still failed to explain how a "distinctive cluster" would survive two thousand years (or even worse ten thousand) of gene flow".
How much gene flow? If they were distinct prior to that gene flow, then the cluster should survive, though detecting it will depend on depth of the cluster analysis, sample size and populations compared.
This would not be the case if there would have been a demic replacement (very distinct from normal gene flow or even occasional minority immigration of some size) but there are no clear grounds to support that demic replacement and in that case, certainly, the cluster would not have survived in any case.
"First of all, your contention that West Eurasians have been diverging for 50 thousand years is (a) irrelevant, (b) unfounded, and (c) probably wrong"...
This is one of our main points of contention and we have discussed on it many different times. It'd be impossible and pointless to start all over.
But even if the divergence of West Eurasians is only from Neolithic times, as seems to be your hypothesis, then we are still looking at one of the main distinctive components of that "Neolithic" scatter. If we exclude Druze and Bedouins (small populations with wacko specificities) it is one of 5 major clusters, all the others being super-regional (widely spread).
And I know of other cases where very small strongly isolated populations show up as distinct even if they could be not so distinct after all. This happened with the Mlabri (an Austroasiatic forager population shown elsewhere as related with other neighbor Austroasiatic) in the HUGO consortium paper on Asian genetics, for instance. So the distinctiveness of the Druze looks a case of that type (not totally sure about Bedouins, it's possible that their distinctive component has a Harifian origin).
"Yep, anyone can understand that your "reasoning" is filled with hand-waiving, special pleading, and that it is motivated by your politics".
What about your politics, Dienekes? Ahem!
My position in favor of native national rights of Palestinians is unaffected by their ethnic or genetic origins. Even if modern Jews would be pure descendants of the Jews of old, which doesn't seem to be the case at all, I would still deny them their alleged right to rob the country of others after 2000 years of exile.
It'd be like claiming half of European Russia for the Turks or the right of Armenians to conquer NE Anatolia (or Cherokee to Appalachian lands) in the year 4000 (now they may still have some claim, as barely a century has passed since the respective genocides).
Whatever the case, I see a clear outstanding Palestinian genetic uniqueness that speaks volumes. Yo prefer to stay deaf? Your problem.
Belonging to the Hellenistic states, the Roman Empire and then Islam makes this period pretty much cosmopolitan quite obviously. This is probably not a particularlirity of Palestine but also applies to that country.
Palestine belonged to "cosmopolitan" empires before the arrival of the Greeks (the Persian one most recently). Thus, your contention that it has been unusually cosmopolitan in the last two thousand years is wrong.
Indeed, with the exception of the periods of Hebrew national autonomy and the period in which Philistines arrived at the coast, it would be quite difficult to find any period of the history of Palestine where the place had a distinctive population element in it.
How much gene flow? If they were distinct prior to that gene flow, then the cluster should survive, though detecting it will depend on depth of the cluster analysis, sample size and populations compared.
Clusters form in periods of relative isolation and low gene flow and are dissolved in periods of unimpeded gene flow.
I leave it to the reader to figure out if Palestine spent any of its time since the Neolithic (with the exceptions I note) in a situation of of impeded gene flow.
But even if the divergence of West Eurasians is only from Neolithic times, as seems to be your hypothesis, then we are still looking at one of the main distinctive components of that "Neolithic" scatter. If we exclude Druze and Bedouins (small populations with wacko specificities) it is one of 5 major clusters, all the others being super-regional (widely spread).
Right, Palestinians are cosmopolitan, they don't have "wacko specificities" and yet they form their own cluster.
I repeat: why do Palestinians form their own cluster if they're like everybody else. You have to come up with an explanation for why there exists a Palestinian cluster. Appeals to Palestinian uniqueness don't count.
My explanation is that Palestinians like these other populations (Mozabites, Druze, Bedouins) are consanguineous Muslims.
What's YOUR explanation for Palestinian specificity. Don't give me any of that "it can't be recent so it must be old BS", tell me exactly when a Palestinian-centered cluster developed and what makes Palestinians special that they should have their own cluster while all these other West Eurasian populations do not.
My position in favor of native national rights of Palestinians is unaffected by their ethnic or genetic origins. Even if modern Jews would be pure descendants of the Jews of old, which doesn't seem to be the case at all, I would still deny them their alleged right to rob the country of others after 2000 years of exile.
Britons got the land from Ottomans, who got it from Arabs, who got it from Romans, who got it from Greeks, who got it from Persians, who got it from Assyrians, who ...
I see no reason to give priority of ownership to Arabs or to anyone else. The Jews don't have possession of Israel because they lived there 2,000 years ago, but because they have the guns and the will to keep it, just as the Arabs had the guns and the will to keep it in the past.
According to your wording, Dienekes, there could never be any sort of differential population clusters because in all cases there is "unimpeded flow" of some sort or dimensions. I bet even Druzes have got unimpeded flow at some minimal levels.
The question is how many actual people immigrates in proportion to natives, normally, even if immigration is "unimpeded", this proportion is low because the country is already settled for the most part.
There may be moments when there is accelerated immigration and even native decline in population (genocide, huge epidemics) but these mostly we seem able to identify only when highly developed industrial or proto-industrial societies have clashed with people with huntergatherer or semi-neolithic lifestyles. Even in a case like Mexico, we still find that people is still some 50% or more of native blood and this native component can be identified by cluster analysis.
"My explanation is that Palestinians like these other populations (Mozabites, Druze, Bedouins) are consanguineous Muslims".
There's no comparison: all your examples refer to small isolated populations (and in the case of Mozabites their specific component seems to be part of a wider North African or Berber component). We see nothing of that in other normal Muslim populations such as Jordanians, Lebanese, peninsular Arabs, Moroccans, Egyptians. Palestinians are just like these.
"What's YOUR explanation for Palestinian specificity".
Palestine is a distinct archaeological region until at least PPNB and so far, in West Eurasia, such clusters tend to be coincident with pre-Neolithic archaeological provinces. The same happens in Europe (Bauchet 2007) where Irish and Poles cluster together, as one would expect from peoples originating essentially in the Rhine-Danube region but Basques cluster differently (Franco-Cantabrian region) and so do Valencians (Iberian region).
That's my most parsimonious interpretation. However, if you don't like it, you may need to look for another period of differential isolation such as the Jewish states of the Iron Age.
"... what makes Palestinians special that they should have their own cluster while all these other West Eurasian populations do not".
It surprised me too. I guess, I strongly suspect, I'm quite sure, that if the the Admixture run would have been allowed to continue to k=16 or whatever, we'd witness the appearance of other such clusters in other populations (some already identified in other studies, some maybe not).
I think that the rather large Palestinian sample, along with some clear old distinctiveness, explains why they popped up as different at such shallow levels.
"Britons got the land from Ottomans, who got it from Arabs, who got it from Romans, who got it from Greeks, who got it from Persians, who got it from Assyrians, who ..."
Right of conquest then? I'm for a more democratic approach: human rights and right of self-determination, sincerely. Violence only brings more violence.
Palestine is a distinct archaeological region until at least PPNB and so far, in West Eurasia, such clusters tend to be coincident with pre-Neolithic archaeological provinces. The same happens in Europe (Bauchet 2007) where Irish and Poles cluster together, as one would expect from peoples originating essentially in the Rhine-Danube region but Basques cluster differently (Franco-Cantabrian region) and so do Valencians (Iberian region).
That's my most parsimonious interpretation. However, if you don't like it, you may need to look for another period of differential isolation such as the Jewish states of the Iron Age.
Ok, so your "most parsimonious explanation" involves formation of Palestinian genetic distinctiveness in PPNB and then its maintenance over ten thousand years despite the fact that Palestine is embedded in a much larger geographical region, and had no noticeable cultural differences from the wider region for most of those ten thousand years.
It's the genetic equivalent of building a sand castle, going back after a month and expecting it to still be there.
It surprised me too. I guess, I strongly suspect, I'm quite sure, that if the the Admixture run would have been allowed to continue to k=16 or whatever, we'd witness the appearance of other such clusters in other populations (some already identified in other studies, some maybe not).
I'm not interested in your speculations. You should just say: "I have no explanation why the only samples that pop up as distinct throughout Eurasia are those of consanguineous Muslims, but I'll pretend that it has something to do with Paleolithic Palestinians".
onur, if you don't find a solution to your "multiple postings of the same comment" problem, I will start deleting all your comments. I'm done cleaning up after you.
onur:
The following sentences of the same paragraph say similar things:
"The genetic closeness, in classical protein markers, of Bedouin to Yemenis and Saudis (Cavalli-Sforza et al. 1994) supports an Arabian origin of the Bedouin. The alternative explanation for the distribution of the Arab-specific haplotypes (i.e., random genetic drift) is unlikely. It is difficult to imagine that the different populations in the Yemen and the southern Levant, in which Arab-specific chromosomes have been detected at moderate-to-high frequencies, would have drifted in the same direction." [emphases mine]
I'm not sure what you're point is. In the earlier paper that discovered the Arab clade, they asserted genetic drift as a possible explanation. In this paper they saying genetic drift is an unlikely explanation. So...?
maju:
"You have totally misunderstood what I said.
I can't but agree that some Arabian Muslims (and surely Arabized Muslims from elsewhere, like Syria, Egypt or Iraq) settled in Palestine. What I say is that there are no grounds and seems most unlikely that there was a massive settlement, much less a displacement of the original population in the Muslim period. "
Let me quote you: "Arabs expanded as aristocratic conquerors and religious missionaries, not as settlers." A missionary non-settler doesn't settle. This is a claim that can't be qualified, only retracted.
"As you insist that I mention my sources, F. Josephus, the main (only?) original source on this matter, talks of 1.1 million killed in Jerusalem alone. However it's very likely that the countryside was largely spared."
I'm very proud of you for acknowledging the importance of historical substantiation. The next step is to take in a balanced approach to the historical material regarding Jewish presence in Palestine in other areas--the Galilee, Tiberias, Hebron--and in subsequent years (notably, at the time of the Islamic expansion)
"I'll take a look to Thomas 2000 (if open access) but, from your quote, it seems to me that he's not talking of the bulk of their haplotype diversity but to a fraction of it (29%) which may indeed represent some historical expansion (not necessarily the Arab-Muslim one) with some founder effects. An Y-DNA expansion/founder effect affecting 29% of the Y-DNA pool may well mean just some 10% or even less of the whole gene pool, considering the peculiarities of gender-bias in demographic processes."
Now let's rewind to quote you again: "Arabs (a bunch of nomads from the desert originally) could not alter the genetic landscape in any meaningful manner." A 29% Arab clade constitutes a meaningful genetic alteration. It directly contradicts your claim of no alteration. And the authors believe this clade derives from the original Islamic expansion, so we're bracketing the question of 13 centuries of subsequent low level clan immigration and bedouin sedentarization. My original assertion was that the PA population cannot be treated as a control sample for "Palestinness" and this is sufficient to show I'm right. The genetics of PA samples is one evidence amongst many factors to be weighed against each other--in light of the historical data--but definitely not to be treated as a control sample. I even somewhat satirically stated, "Your treatment of the modern Palestinian Arab population as a millennia-old Palestinian genetic constant seems to be an assumption or axiom on your part"--and your response was to completely agree to this characterization! I'm sorry, but "clownish" comes to mind.
onur:
Insultation? haha that's worse than impudent :P
"History is essentially nothing but a discipline of interpretation of historical sources and artifacts as realistically as possible, and like in many things open to interpretation there are many different and often conflicting interpretations in history."
None of this contradicts what I have said. That you would think so suggests you've perhaps contorted the meaning of what I've said far outside the intended meaning due to the rigidity and crudeness of your way of thinking ;)
"For instance, the existence of the Roman Empire is one of those many undeniable truths"
So you agree that there is a core confluence of historical opinion built up over space and time. Next step is to investigate how large that core is, including its very large perimeter consisting of reasonably conflicting debated interpretations. Then we look at the surrounding penumbra of intelligent historical revisionism, and after that we can see the linings of unintelligent revisionism.
"We also know from primary historical sources and artifacts that some Arabians migrated to Palestine"
Maju didn't seem to know this. I'm glad I'm not the only one telling him now. He specifically denied any Arabian settlement in Palestine, although in his last posting he apparently has tacitly retracted the claim.
"Arabian-originated Muslim migrations might have made a major impact on the genetic landscape of Palestine, but historical sources aren't and cannot be helpful for us in learning the proportions of this impact"
Well they can, they just can't be definitive. However it is unhelpful if they're dismissed on grounds of personal political preference rather than the use of countering historical sources (which is what maju has been doing).
Kidproquo: I'm trying to disengage from the discussion, because it's already long enough and some parts of it have not been very nice. But as you insist...
"Let me quote you: "Arabs expanded as aristocratic conquerors and religious missionaries, not as settlers." A missionary non-settler doesn't settle. This is a claim that can't be qualified, only retracted".
It can perfectly be qualified. That's the essence of Arab-Muslim expansion in the times of the first Caliphate. I'm retracting nothing of this.
If some individuals or groups also settled here or there, that's the exception, not the rule.
Furthermore Arabia peninsula did not have the demographics to colonize, even if they wanted to, in meaningful numbers in the vastness of their conquests. Not as to significantly alter the local demography.
They are also not known to have caused any democide, which could help in such demographic alteration you claim. It was essentially a military, political and religious conquest.
However, in the course of ulterior history, these areas, now rather unified culturally (and sometimes also politically), were in mutual relation of certain cosmopolitanism, so it's possible that in the period after the Arab-Muslim conquest till present day, some localized colonizations may have happened too.
This is my qualification. And I don't care if you accept it or not. Too tired of this discussion.
"I'm very proud of you"...
Thanks daddy. O_O :?
"The next step is"...
That you acknowledge that the situation upon the Great Jewish Revolt was much more susceptible of demic change, than any moment since the Muslim conquest till 1945.
By that, I don't mean that there was replacement. I don't think so. Just that the opportunity for such a radical demic change as the one you propose is much more clearly available than at any other historical moment before the arrival of the Zionist settler-conquerors.
"Now let's rewind to quote you again"...
Rewind to your mamma. :(
maju:
Disengage away, however I'll continue to respond to what's addressed to me.
"If some individuals or groups also settled here or there, that's the exception, not the rule. "
Given the entire point of the discussion is the impact of such settlement, not just during the first caliphate but during the whole of the Islamic period, your attempted exception is exceptionally disingenuous
"They are also not known to have caused any democide"
No such claim was made. However I notice you tend to avoid the issue by attempting to limit the period of discussion to a handful of years.
The claim that was made was that the impact of Muslim immigration and bedouin sedentarization over 13 centuries has been substantial enough such that the PA population cannot constitute a genetic control group, that there has been a meaningful genetic alteration of the Palestinian landscape. Actually I was caught off guard that you were unaware of the Arab clade--I naively assumed you were familiar with the breadth of the genetic literature on the matter. In any event, the 29% (actually higher in the previous study) Arab clade is sufficient to put the matter to rest, and the rest of the genetic evidence previously discussed only adds to the substantiality of the impact.
"because they have the guns and the will to keep it, just as the Arabs had the guns and the will to keep it in the past".
And using that as a justification surely denies the Israelis the right to adopt the high ground when Palestinians fight back using suicide bombers. It's just that they don't have sufficient guns.
Are you Jewish, Mr Dienekes?
Ethiopian and Ashkenazim are both true jews in my hypothesis, the "L2" mtDNA marker is present in the two populations also the derived and sibling mtDNA Hg "M" and "N", as well as the Y markers Hg E3b and 4s too, all of this from East Africa and so. They belong respectively at one of the three nucleous or center jewish ancient populations, that evolving the called "Syrian-European nucleous"(helenistic and Roman times).
The oldest center -Ethiopians belong these- were that developed in Napata and Elephantine (Kush) and whose nucleous or center was after Alexandria, and I called "Coptic Nucleous" derived in two bias, and split forwards the North via Europe -intermixed with the Syrian Europe nucleous- or the South, via Nile and the Horn Of Africa.
The "Babilonian and Persian nucleous" is other of the above three mentioned centers and included Bukara, Iranian and Iraki mainly.
All of this Nucleous take Judaea and Israel like a axis and pendulo.
Another fourth Nucleous or center I call "East Europe" -not mainly conected with ME-, is not ancient like the three others and was the Jewish Khazar Empire stiring into Askenazy current population and others. All of this events were naturaly intrajewish asimilations in all jews current populations.
The Ashkenazim hyperhaploydia is explained by the superposition and overlay of diverse fount or source population , that are all of this of Jewish origin (that consider converted into intraJewish assimilations) , one coming from the “Syrian European nucleous” – that Sephardic as well as preAshenazim bring inside -. The other convergence were the “Coptic Jewish nucleous”, coming from Alexandria, the main and largest Judaic center in ancient times – the buried and graves in Jewish graveyards and catacombs of Tuscan, and Alsace as too Rhineland cities take a lot of Egyptian ornaments and display figures from these, as well as Y and mtDNA markers - . The great Jews migration from Egypt beginning after the Muslim invaders from Arabia in the VII AE century. The “Babylonian and Persian nucleous” take place and contacts newly with and when the “preAshenazim second fase” were migrating to the East Europe. A remarkable contact was with the fourth “East Europe Jews nucleous”-not related or little related with ME-, with the descendant of the Jews Khazarians ones, spreading every where and carrying a lot of East Europe and Eurasian markers. That happen between the XI and XII century AE.
The Tuscan host populations come from Anatolia like infers mtDNA markers, and others, yet present today – a thread Etruscan link - and are so common in South East Basin like Albanian, Grecian, Tunisian and Anatolian , as well as the entirely Italy and some South France spots, practical absent in central or North Europa or East Asia. If we compare Ashkenazy jews with this South European Tuscan population will see a more European genes pool coming from Europe than if we compare with Central and North European population.
Dr Hector H. Otero C.
Argentina.
See too:
http://www.nature.com/news/2010/100603/full/news.2010.277.html
See, Comment 11149 and 12952 with table 1 -partial-, remember that the sibling Hg "M" and "N" -from "L3"- too correspond to East African origin, and are not included at all-see complete table 1, in reference-.
www.scirp.org/journal/PaperInformation.aspx?paperID=19567
Post a Comment