 This is a nice paper on European population structure with a large number of SNPs, which comes at the heels of some earlier studies:
This is a nice paper on European population structure with a large number of SNPs, which comes at the heels of some earlier studies:- Genetic structure in Northern Europe with 250K SNPs
- Geography and Genetic structure in Europe (again)
- 500K SNP Europe-wide study of genetic structure
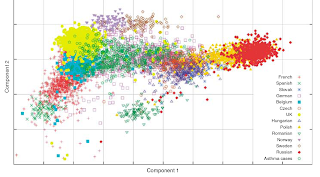
- In this study the first principal component of variation is along an east-west axis, rather than north-south as in previous studies. This is due to the limited number of southern European populations, and the great number of populations along an east-west axis from Spain to Russia. As I have mentioned before, the results of a principal components analysis are dataset-dependent.
- The nice technique of this paper is to infer the ancestry of an unknown sample (which could be perhaps a forensic case or customer of an ancestry analysis test) using only summary statistics. Imagine that you have 1,000 individuals from different populations, and want to guess the ancestry of an unknown test case. You could go about doing a full STRUCTURE run using the 1,001 individuals, or you could exploit the information garnered from an analysis of the 1,000 individuals (a PCA analysis in this case) to test the 1,001th individual. This is much faster and convenient, since the full STRUCTURE run is very time consuming.
Some ethnic groups are clearly distinguishable from each other (e.g. Swedes vs. Spaniards); some groups are partitioned into fairly disjoint sets (Spain I vs. Catalans in Spain II); others mutually overlap (e.g., British and Irish); while others overlap asymetrically (e.g., some former Yugoslavs in the Greek cluster, but not vice versa).In this paper, the authors did a systematic study of the "ethnic distinctiveness" of their samples. In a first experiment, they used 80% of their data to identify the features of the various populations (e.g., Germans or Spaniards), and then tried to guess the origin of the remaining 20%:
 It is clear that some nations appear to be distinct. For example, most test Spaniards (94.5%) are correctly guessed as Spaniards, with some (5.5%) guessed as French. Of course, this distinctiveness would be reduced if further populations (e.g. the Portuguese) were added to the analysis. More strongly, 99.1% of Norwegians are guessed correctly as Norwegians.
It is clear that some nations appear to be distinct. For example, most test Spaniards (94.5%) are correctly guessed as Spaniards, with some (5.5%) guessed as French. Of course, this distinctiveness would be reduced if further populations (e.g. the Portuguese) were added to the analysis. More strongly, 99.1% of Norwegians are guessed correctly as Norwegians.Other nations appear to be less distinct. For example, only 45.3% of Slovaks are guessed as Slovaks with most of the remaining ones guessed as Czechs (25%) or Hungarians (22%).
In some cases there is asymmetry of affiliation. For example, no Belgians are guessed as Germans but 10.2% of Germans are guessed as Belgians. Similarly 9.9% of Swedes are guessed as Norwegians, but only 1% of Norwegians are guessed as Swedes. While each case needs to be addressed individually, this observation is consistent with historical asymmetry in immigration patterns or ethnic identity formation. So, while e.g., the bulk of Germans (64.4%) are guessed correctly, sizeable minorities are guessed as Czechs, Belgians, or Scandinavians.
I would speculate that large central European countries have historically (both due to prestige or geographical position) absorbed more diverse populations from neighboring nations, while smaller peripheral countries have mostly acted as sources of population, reserving their own genetic distinctiveness.
In a second experiment the authors guessed the origin of individuals, but excluding the country from which they actually originated.
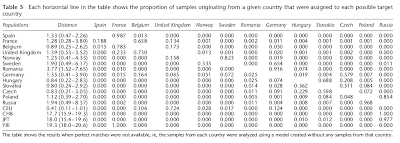 Once again, it is clear that members of particular nations can mostly be mistaken for members of their closest neighbors. Almost all Spaniards are guessed as French; French mostly as Belgians but with sizable Spanish and UK minorities; UK as Belgians but with sizable French minorities; Norwegians mostly as Swedes but with some UK; Swedes mainly as Germans but many as Norwegians; most Poles as Russians, but some Slovaks or Czechs, and so on.
Once again, it is clear that members of particular nations can mostly be mistaken for members of their closest neighbors. Almost all Spaniards are guessed as French; French mostly as Belgians but with sizable Spanish and UK minorities; UK as Belgians but with sizable French minorities; Norwegians mostly as Swedes but with some UK; Swedes mainly as Germans but many as Norwegians; most Poles as Russians, but some Slovaks or Czechs, and so on.The importance of these results can't be underestimated. While it can be argued that some ethnic groups are spuriously distinctive only due to insufficient sampling of the geographical continuum, it is more difficult to do this for others. For example, it is now possible to identify particular ethnic groups, e.g., Norwegians, with great accuracy from DNA.
More markers and more populations will doubtlessly enhance our ability to distinguish European nations using DNA. But perfect accuracy is unlikely; in most European nations there are probably minorities which -for historical reasons- allied themselves with one country or political entity even though they were ultimately of different genetic background than the majority population of that entity.
Nonetheless, at a time when -due to a sort of mental hysteresis- proclamations that "races are social constructs" are still routinely made, the discovery that not only races, but even closely related ethnic groups (e.g. Norwegians and Swedes) can be distinguished with greater than 90% accuracy, serves to illustrate the scientific irrelevance of the ethnic nihilists and the affirmation that nations are, at least in part, genetic entities.
European Journal of Human Genetics (2008) 16, 1413–1429; doi:10.1038/ejhg.2008.210
Investigation of the fine structure of European populations with applications to disease association studies
Simon C Heath et al.
Abstract
An investigation into fine-scale European population structure was carried out using high-density genetic variation on nearly 6000 individuals originating from across Europe. The individuals were collected as control samples and were genotyped with more than 300 000 SNPs in genome-wide association studies using the Illumina Infinium platform. A major East–West gradient from Russian (Moscow) samples to Spanish samples was identified as the first principal component (PC) of the genetic diversity. The second PC identified a North–South gradient from Norway and Sweden to Romania and Spain. Variation of frequencies at markers in three separate genomic regions, surrounding LCT, HLA and HERC2, were strongly associated with this gradient. The next 18 PCs also accounted for a significant proportion of genetic diversity observed in the sample. We present a method to predict the ethnic origin of samples by comparing the sample genotypes with those from a reference set of samples of known origin. These predictions can be performed using just summary information on the known samples, and individual genotype data are not required. We discuss issues raised by these data and analyses for association studies including the matching of case-only cohorts to appropriate pre-collected control samples for genome-wide association studies.
Link

96 comments:
Interesting, is there a lot've asthma among Poles and Russians? Not really sure why that is present here, as seems to clutter up a lot've the chart in the center there-- but I didnt read the entire study yet.
Well this is interesting aside from the Geographical stuff which is to be expected.. the Hungarians are strongly seemingly linked to the Poles and Russians, which could be the strong "Neo-Danubian/Uralic" connection among them. Romanians are closer to South Western Europeans..some German and a few Polish/Russian samples seem to headed that way too.
The Belgians being closer the UK, might be more in synch with history, as its supposed to be the Balgae which the Briton's got most of their culture/blood from, not the Celts(Gauls) proper during the Bronze or Iron Age.
^ Crimson you need your head checked.
Dienekes wrote,
"Nonetheless, at a time when -due to a sort of mental hysteresis- proclamations that "races are social constructs" are still routinely made, the discovery that not only races, but even closely related ethnic groups (e.g. Norwegians and Swedes) can be distinguished with greater than 90% accuracy, serves to illustrate the scientific irrelevance of the ethnic nihilists and the affirmation that nations are, at least in part, genetic entities."
You missed the more important point: It means Norwegians and Swedes are separate races, which makes the concept of race again artificial, since it appears to be merely geographical clines.
There is a sizeable population of Hungarian speaker ethnic Hungarians in Slovakia, so the identification is correct.
On the other hand, the Roma are no less numerous but they dont appear on the study.
which makes the concept of race again artificial, since it appears to be merely geographical clines
It's a bit more than that with Norway, which had much more settlement based on fishing than Sweden, but in general, I agree.
We are just a couple of years away from basically distinguishing what is left over and commingled from extended families of the agricultural expansion. That's not races, that's cousins who outmarried.
But perfect accuracy is unlikely; in most European nations there are probably minorities which -for historical reasons- allied themselves with one country or political entity even though they were ultimately of different genetic background than the majority population of that entity.
I agree that perfect accuracy is unlikely, impossible I'd say. But I tend to disagree with the rest of your claim: you are basically trying to ignore the admixture (and internal sorting) event that happens with every sexual reproduction in every single generation. The elements from neighbour countries are not due to "minorities" but to normal genetic flow across ethnic and political borders (as well as the flow of such borders themselves). There are no absolute barriers to admixture between any two populations, except total isolation and (even for the case of remote Norwegians) this total isolation has never existed within Europe, much less if we consider invasions and migrations of all sorts through history and prehistory.
Populations, specially when isolated, do tend to become more or less homogeneous but this process is also breached all the time. So perfect identification will never be possible unless you get the people and put them in neatly separate boxes for many generations, what in itself is another impossible thing to do (much less desirable).
---
As a side comment, it is certainly interesting how the E-W cline is emphasized in this particular study, once excluded most SE European populations. Unlike what Dienekes says, not all previous studies had emphasized the N-S axis, some had also noticed the E-W one, even as dominant. This may depend largely on the sample sizes and choices, because it's impossible to depict such multidimensional variability in a bidimensional plot.
You missed the more important point: It means Norwegians and Swedes are separate races, which makes the concept of race again artificial, since it appears to be merely geographical clines.
Race classification is hierarchical. Just as Homo sapiens can be subdivided into races, so can the Caucasoid race be further subdivided, and this structure will probably go even deeper once large-scale country-wide samples are assembled.
The elements from neighbour countries are not due to "minorities" but to normal genetic flow across ethnic and political borders
Geography is not sufficient. Poland may be geographically closer to Germany than to Moscow, but it is much closer to the latter genetically. Spain is closer to Romania than to any of the intervening countries (save Romance-speaking France and Belgium).
Btw, there are actually two seperate German samples here, and the Dresden one, originating in former West Slavic territory, does overlap with the Lodz Poles.
You are looking only at the scatterplots of the principal components. What seems to overlap in one PC may in fact be non-overlapping in higher order PCs. And indeed, from Table 4 it is clear that 0.2% of Poles were assigned to Germany and 0.8% of Germans as Poles. And in Table 5, 0.7% Germans are assigned to Poland and 0.1% of Poles to Germany.
So, the two nations seem to be almost perfectly distinguishable from each other, with almost no mistaking a Pole for a German or vice versa.
Ren:
The fact that Norwegians and Swedes can be distinguished genetically does not mean that race doesn't exist. What it means is that ethnicity, like race, is a biologically reality and not a social construct.
SNPs don't indicate a genetic basis of race - they indicate genetic isolation over a potentially short period of evolutionary time. There are obviously some minor morphological, pigmentation and (rarely) disease susceptibility differences among ethnicities, but any loci that contribute to traits that really matter (and are generally the focus of racial steroptypes)e.g. morality, work ethic, sexuality, etc. certainly will vary more within groups than among them.
Geography is not sufficient. Poland may be geographically closer to Germany than to Moscow, but it is much closer to the latter genetically. Spain is closer to Romania than to any of the intervening countries (save Romance-speaking France and Belgium).
Ahem! Spain is not closer to Rumania than to, say, Norway in that graph. It is certainly closer to "Germanic" countries like Germany or England.
The case of Poland may well have been radically distorted by recent (post-WWII) mass demographic movements that brought Eastern Poles to former German provinces. It was surely not that way at the beginning of the 20th century, wen the cline was with all likehood much smoother.
...
SNPs don't indicate a genetic basis of race - they indicate genetic isolation over a potentially short period of evolutionary time.
Strongly agree with this observation: (relative) inbreeding certainly should account for spontaneous statistical homogeneity.
I would add a comment of the sort of "what the heck is a French?" Someone from Champagne (probably close to Belgians and Renanians), from Provence (surely close to North Italians or Catalans), Gascony (Romanized Basques) or what? While smaller nations are surely more homogeneous, large nations such as France, Spain, Germany, Italy or Russia certainly deserve a regionalized approach to account for their obvious plurality, specially in historical and therefore genetic terms.
maju said,
"I would add a comment of the sort of "what the heck is a French?" Someone from Champagne (probably close to Belgians and Renanians), from Provence (surely close to North Italians or Catalans), Gascony (Romanized Basques) or what? While smaller nations are surely more homogeneous, large nations such as France, Spain, Germany, Italy or Russia certainly deserve a regionalized approach to account for their obvious plurality, specially in historical and therefore genetic terms."
Bravo! I've been trying to say the same thing for a long time. What is a 100% French person? At what point in time did anyone become 100% French? Is a Parisian more French than a Lyonnais? If the genetic constitution of present individuals is due to geography's constraint on the mating patterns of previous generations, then one should be able to find geographically distributed genetic variation within the population of a single country of the same sort (though probably smaller on an absolute scale) as that which many researchers have found among the populations of several countries.
MAT:
If SNPS don't indicate a genetic basis for race then what does? The more that I read and listen to the advocates of the social construct theory of race the more they seem like left-wing versions of creationists who just can't accept the scientific evidence because it disagrees with their personal views.
Spain is not closer to Rumania than to, say, Norway in that graph. It is certainly closer to "Germanic" countries like Germany or England.
Fst(Spain,UK)= 0.0024
Fst(Spain,Norway) = 0.0047
Fst(Spain,Romania) = 0.0023
Dienekes,
Let's sample more East Germans (especialy former Slavs from Rostock), and then also Silesians, Pomeranians, etc.
Then we'll talk...
In fact..
Why should Poles be mistaken for a lumped German sample including Munich??
Munich isn't anywhere near Poland. And it never had any Slavic roots.
It was more Celtic than anything.
Btw, how come no one is discussing the African and Asian affinities here??
Brits have the least, followed by Norwegians and then Poles.
Germans are way down the line. And Romanians are near the bottom. Wonder why that is?
"If SNPS don't indicate a genetic basis for race then what does? The more that I read and listen to the advocates of the social construct theory of race the more they seem like left-wing versions of creationists who just can't accept the scientific evidence because it disagrees with their personal views."
That is exactly the way I see it. I never believed them to begin with but I often cannot get over how many recent biology and other science graduates believe race is only a "construct", and biologically meaningless.
This information is very fascinating but mostly not surprising except for the asymmetries. No amount of information will ever convince the race-denialists though, since they don't care a whole lot for real science considering that their beliefs are more like a religious faith based on political correctness.
I'm not familiar with the 'social construct theory of race,' but I'll check it out. From what I know, humans have only had a few tens of thousands of years to diverge across the planet, which isn't really enough time for much genetic change to occur.
The thing with the whole concept of "race" is that if it just refers to the color of your skin and the shape of your nose, it's pretty trivial. It's only when "race" is thought to account for differences in morality, intelligence, etc. that it matters if race has a genetic basis. This type of racial difference is incredibly unlikely since humans have very little genetic variability in the first place, we diverged a very short time ago, and we've all been hunter gatherers living with similar selective pressure for most of the time since.
This will all be moot in a few more years when we have thousands of whole genome sequences from different individuals...
Average Joe wrote,
"The fact that Norwegians and Swedes can be distinguished genetically does not mean that race doesn't exist. What it means is that ethnicity, like race, is a biologically reality and not a social construct."
I don't think you understand what I'm saying. Let me put it in a less abstract way so you can understand.
What makes Norwegians and Swedes not separate races if they can be distinguished genetically?
Fst(Spain,UK)= 0.0024
Fst(Spain,Norway) = 0.0047
Fst(Spain,Romania) = 0.0023
But not in the graph, where the centroid of Spain is like 4+ grid units from the Rumanian one (along the PC1), only 2 from the UK (mostly along the PC2 axis) and about 3.5 from Norway (again mostly along the PC2 axis).
Rumania and Spain may both have a language derived from Latin but had no historical relation whatsoever (apart of those that can be generalized to all Europe). Jamaica has the same language as Canada and well, I would not expect too much genetic connection honestly. Language and genetics are different things.
Also the most closely related to Rumanians by Fst stats are (in this order) Hungary, Slovakia, Germany and Czech Republic (tied), France, Belgium (and only then Spain much behind in the figures, almost as distant as the UK). It doesn't say much in favor of any particular "Latin" or "southern" connection either.
It's curious though that by Fst Rumanians appear pretty much distant from Poles and Russians (more than to Spain and UK) what makes them appear somewhat "western" in the whole context (also reflected in the graph). It's quite odd, because their closest Fst relatives, Hungarians, appear much more "eastern" (a lot closer to Russians, for example - and this is also reflected in the graph).
Btw, how come no one is discussing the African and Asian affinities here??
Because they are very remote. If anything fig. 2a shows clear is that (with the exception of a single Japanese dot) there are no real intermediates.
Brits have the least, followed by Norwegians and then Poles.
Not really so. There are some diverse scattered samples that tend more to the "south" than the main cluster (fig 2b, same criteria: comparison along intercontinental PCs - do not use the other Euro-only plots for that, please). If you draw an imaginary line towards the YRI (Nigerian) sample, the closest ethnic cluster are Spaniards but the closest individual samples are French and British (some Belgians and Germans too). If you do the same towards the Japanese/Chinese cluster, the nation with a more marked "eastward" tendency are Russians clearly but also do some Rumanians, Germans, Brits and single Slovak and Swedish individuals.
And for those who just want to argue about the reality of races, I must say that the issue is not wether there are some genetic or phenotype differences but wether such things really make any difference at all.
We are always, in these issues, selecting the small differences and ignoring the huge similitudes and that is because we want to know about the fine detail of Humankind. And that is good because we are a very curious species and we like to research and find out. But we cannot forget that the similitudes are infinitely much larger than any differences we can spot. Let not the trees impede the vision of the forest, ok?
I'm not familiar with the 'social construct theory of race...
It's as simple as to say that Obama is "black". That is a clear example of social construct because he's actually (biologically) as black (Tropical African) as he's white (Creole European).
Basically the modern concept of race is largely a socio-cultural construct. While Romans had contact with Nubians for instance they did not have a theory of races like we do now and their use of the word was more like "Norwegians and Swedes": stocks n general. They did not concieve a "white race" or a "black race", even if they knew representativs of them.
We reinvented the concept of race in the age of exploration and colonialism in line with socio-economical and cultural factors, with slavery and other types of colonialist domination playing an important role in it. Significativley at the beginning of that period religion was the important factor when determining who was morally entitled to be opressed (hence most Africans could be enslaved because hey were not Christian but Christian Africans were given a muchmore horizontal treatment, same among Muslims - even the Irish were enslaved by the English on religious grounds, specially under Cromwell). But gradually the issue was transformed in one of mere mega-stocks (races as we know them now) as the role of the different colonized/enslaved peoples became more and more specific.
So from ethno-religious discrimination we arrived to racism. And racialist emphasis is often just undercover racism, to be clear.
Maju,
Look at table 1. Add up the last three values.
1. Brits
2. Norwegians
3. Poles
However, some individual Brits do show extra affinity to Asia and/or Africa.
Look at table 1. Add up the last three values.
You cannot add up East Asians and Tropical Africans like that. You must consider the affinities with the two extra-European samples separately.
In general it's interesting that Europeans (all Europeans) appear markedly closer to Tropical Africans (almost double values) than East Asians do. Though logicaly we are closer to East Asians than to Tropical Africans.
It's also interesting that Chinese and Japanese samples are more Fst distant among them than any two European samples whatsoever (though Eastern Europeans and Spaniards approach that mark).
However, some individual Brits do show extra affinity to Asia and/or Africa.
That's pretty obvious and not just Brits. Some of those individuals appear more North African than properly European.
But you should measure African and East Asian affinities separately in any case. They may add up though: I imagine that West Asian influence (naturally more intense in the Balcans) could bridge the gap somewhat towards both (always within West Eurasian parameters anyhow).
Erratum: not "almost double", just like 25% closer to Tropical Africans than East Asians are (messed up the figures, sorry).
Yeah you can add them up roughly, for that prupose.
Basically, the higher on graph 2, the more European. So it works just fine.
It's curious though that by Fst Rumanians appear pretty much distant from Poles and Russians (more than to Spain and UK) what makes them appear somewhat "western" in the whole context (also reflected in the graph).
I think there is a lot of recent thought that like (at least the cis-alpine) Galliae, people in the region of Romania spoke a - dare I say it - "Celtic" language with close affinity to Latin, which made it so easy for them to become romanized.
Since I abhor that oversubscribed misnomer, I will just say that clearly, many tribes around the alps 2000 years ago spoke languages that were at least partially mutually comprehensible. However, when I try to read the few "Celtic" words left, I can honestly not say that I find them any closer to Latin than to old German... (hence my criticism to associate anything and everything "Celtic" with either Latin-like or -worse- Gaelic languages).
At any rate, people living in the region of Romania 2000 years ago were linguistically, and by proxy genetically, closer to other folks surrounding the Alps. It may very well be that even until today, they have been less impacted by Slavic expansion than surrounding regions.
But not in the graph, where the centroid of Spain is like 4+ grid units from the Rumanian one (along the PC1), only 2 from the UK (mostly along the PC2 axis) and about 3.5 from Norway (again mostly along the PC2 axis).
PC1 and PC2 are by definition a part of the overall variation, and as I said Spain is closer to Romania than to either UK or Norway.
The distance between Poland and Munich/Dresden is 0.0012.
Anyone know roughly what it would be if Munich was dumped?
I'm wondering how much actual Kelto-Germanic heritage Dresden has (its root is "wood" in Slavic).
The area surrounding the Elbe (and Saale, Spree) rivers were "Kelto-Germanic" (I kind of like that ;)) until about 500 AD, when the Slavic Sorb tribe settled in large numbers in exactly those regions. IMO, they never completely dominated these regions, but had a large presence. Many geographical names of this region, all the way to the Baltic, now have alternating Germanic and Slav roots. However, historically, that is a very recent development, and in later times, the Sorbs (or "Wenden") almost always were a minority on the defense.
In recent (last 500 to 1,000 years) history, culturally, linguistically, and almost certainly also genetically dominantly "Germans" largely again (or always) have outnumbered Sorbs in this region. Many of the classical German writers are from this eastern region of middle/high German, including Martin Luther, and the region has been one of the prime centers of German culture and innovation for much of the past 500 years. (Also, despite its recent massive historical and economic disadvantage, it also scores highest in Germany on the PISA test...).
Of course, on the other hand, much of central east Germany has been R1a and I (and not R1b) for more than 4000 years...
I think there is a lot of recent thought that like (at least the cis-alpine) Galliae, people in the region of Romania spoke a - dare I say it - "Celtic" language with close affinity to Latin, which made it so easy for them to become romanized.
Dacians did not spoke Gallic. The relation of Gaelic with Latin is not straightforward either. The reason why Romania speaks a romance language and not Slavic, Turkic or Hungarian beats me really. It's one of those unimaginable oddities of history, specially if you ponder that Dacia was part of the Roman Empire for a rather short period and had no Catholic influence.
At any rate, people living in the region of Romania 2000 years ago were linguistically, and by proxy genetically, closer to other folks surrounding the Alps.
I have not the slightest idea why do you associate Romania with the Alps and Celts, really.
...
PC1 and PC2 are by definition a part of the overall variation, and as I said Spain is closer to Romania than to either UK or Norway.
Not by PC1 certainly.
...
Anyone know roughly what it would be if Munich was dumped?
There would be two separate clusters: western and eastern. Munich (well, Germans in general) holds them together (see fig 6).
I'm wondering how much actual Kelto-Germanic heritage Dresden has (its root is "wood" in Slavic).
It doesn't really matter, IMO: German colonization (and assimlation) of the area was very intense in the Middle Ages (it had also been Germanic before, Celtic even before, etc.). What matters is that the Dresden sample is between Munich and the Czech Republic (see again fig. 6). Germans (at least in this study) hold Europe together.
Yeah, but just looking at the plots, roughly, would it be 0.0006 or something like that?
But yeah, if Munich was dropped, then the differene between Germans and Czechs/Slovaks would drop to almost 0%.
And Maju, I wouldn' be so dramatic. The reason west and east extremes aren't mixing is because of the Germans. They were in the way...and at the same time, absorbed people from both directions.
Romenians, as the Bulgarian, are for the great part from the ancient Illirian-Trachis people. Them where romanizated easily, because the commercial activity and the military presence were very strong. Roman influence went over the frontier and also the Free Dacians were romanizated. The Germanic and the Slavic pressures and in the Balcans were strong, but a lot of people withdrew him on the mountains to be not absorbed and to maintain their language and their culture. They had called Vlachs from the Slavics but them always called him Romans, Romenians, Arumenians, Istrorumanians. Vlach means "a man speaking a latin language. Poles today call Italy "Wochy", and the Hungarians"Olaszország"yet
The population of Europe in paleolitic age has happened from South toward North.For This Romenians and Bulgarians are closer to the other people of Balcans, Albanians and Greeks and and more close to the other people of the South Europe, as you can see in the previous studies, Slovenians, Croatians and Serbs are very distant from Italians, their neighbors, adjoining instead to the Cechzs and Poles.
The Hungarians are for the most greater part magyarizated Slavs so them are next to the other Slavics. In every way the ancient Pannonia, a plain region and without strong geographical borders it is the most genetically mixed of Europe.
maju said,
"In general it's interesting that Europeans (all Europeans) appear markedly closer to Tropical Africans (almost double values) than East Asians do. Though logicaly we are closer to East Asians than to Tropical Africans.
Erratum: not "almost double", just like 25% closer to Tropical Africans than East Asians are (messed up the figures, sorry)."
I don't see that reflected in the graph of the principal components. The graph has the East Asian cluster being only 2/5 of the distance of the European cluster from the Yorubans along PC1, which describes nearly twice as much of the variance as PC2. The European cluster is equidistant between the Yorubans and the East Asian cluster in regard to (the much less significative) PC2.
In other words, these East Asian samples are closer to this Yoruban sample in regard to PC1, but these European samples are closer to this Yoruban sample in regard to PC2, unless there has been some distortion in the scaling of the graph's axes.
But yeah, if Munich was dropped, then the differene between Germans and Czechs/Slovaks would drop to almost 0%.
Eastern Germans, not "almost zero": continuity and overlap are different concepts.
And Maju, I wouldn' be so dramatic. The reason west and east extremes aren't mixing is because of the Germans. They were in the way...and at the same time, absorbed people from both directions.
That's obvious, man. I just said that if you drop Munich and/or Dresden from that sample the continuum is broken. So, yes: Germans "keep Europe together" at least in what regards to this study. They do not because of being German but because of being Central. But they are the only ones in this study that really save the gap (no Italians sampled).
I don't see that reflected in the graph of the principal components.
Ebizur: I took those figures from the table. The scatterplot only values like 1.59% of the whole matter, so there may be some differences. The scatterplot seems to measure basically along the specific African-European and African-East Asian axis of difference (not much difference by the way when it only ammounts to 1.59%). The European-East Asian differential is not directly measured/plotted there.
First comment by Ren was said beautifully said ,I like your eloquence.
I'm still wondering though ,when are they gonna use this on REAL people to classify one thing or another-And even put it on all their public documents so it'll be like a major fact about you-and you can still pick whatever you like ,but at least you'll know what you're workin' with.
Ren:
The major differences between races and ethnic groups is that ethnic groups refer to subgroups within the same race. For example, Swedes and Norwegians are different ethnic groups who belong to the Caucasoid racial group. Because Swedes and Norwegians come from the same racial group they are going to be more closely related than different racial groups such as Caucasoids and Negroids.
Some websites for those who doubt the biological reality of race:
http://www.rlynn.co.uk
http://psychology.uwo.ca/faculty/rushton_pubs.htm
More on race:
http://armandleroi.com/media/pdf/AML_journ012.pdf
http://www.goodrumj.com/Edwards.pdf
http://evoandproud.blogspot.com/2008/06/lewontins-fallacy.html
http://www.vdare.com/Sailer/sarich_miele.htm
The major differences between races and ethnic groups is that ethnic groups refer to subgroups within the same race.
Not really: Cubans are an ethnic group and belong to different races (as per the usual classification schemes), same for US-Americans or for nearly any ethinicity nowadays. Ethnicity is a matter of self-identification and more or less consensual external classification and is most often than not directly related to language. Ethnicities are socio-cultural and can perfectly be multirracial (though in the long terms they tend to homogeneize their genetic pool).
In fact the term race originally meant ethnicity or stock quite diffusely. There's no valid universal "scientific" terminology in all this: it is certainly a social construct to a great extent. There is some reality in all those differences but it's not something that can be put in neatly packed boxes at all: there's too much subjectivity and cultural components in them.
For example, Swedes and Norwegians are different ethnic groups who belong to the Caucasoid racial group.
Are you telling us that there are no Swedes or Norwegians that have no Tropical African, South or East Asian or Native American blood? I do not know their reality closely but I certainly know of many Basques that are from very different stocks, both from Europe and from out of Europe. A Basque with Chinese blood? Why not? Basque-ness is defined by language and self-identification not genes. In fact they do exist and are more frequent each day. And the same applies elsewhere, with language and that feeling of belonging being nearly everywhere much more important than any phenotypical difference.
...
Some websites for those who doubt the biological reality of race:
http://www.rlynn.co.uk
http://psychology.uwo.ca/faculty/rushton_pubs.htm
LOL, Rushton and Lynn, Lunn and Rushton... are there any less prestigious individuals on this Earth?
Those people are doing pseudo-science and are not better than the parapsychologist of the corner. Though the parapsychologist is surely a much better and respectful person by all measures.
Ignoring white collar neonazis, one can argue that biological race does exist on grounds of geographical phenotype-associated genetic clustering. But that more often than not ignores clines and clines are as important as clusters and one would not exist without the other. If races exist, immense gray zones of mestizage also do exist (and I would say that are prevalent).
No neat boxes for you this Christmas, sorry.
"Ignoring white collar neonazis, one can argue that biological race does exist on grounds of geographical phenotype-associated genetic clustering. But that more often than not ignores clines and clines are as important as clusters and one would not exist without the other. If races exist, immense gray zones of mestizage also do exist (and I would say that are prevalent)".
It would be difficult to explain it better than that.
Sound,
That's not true that Polish and Germans never married - around Poznan they certainly did, and all over Silesia.
I know German-Poles and Polish-Germans.
Many Poles settled in the Ruhr valley in Western Germany after WWII, and are considered German. Whereas many Polanized Belarusians and Ukranians settled in vacated German lands - like Prussia and Silesia - after WWII to become Poles.
Both German national soccer team strikers are Polish.
Copernicus - a Pole from Torun - wrote his great works in German.
These 2 peoples have always had a close relationship, though not always amicable.
Bavarians and Swabians have always been Celtic, they are part of the Celtic homeland. If you know real Bavarians and ask them what ethnicity they are, they will answer, "Bayerishe", not "Deutsche"!
I see "Sound of the Occident" has lost his marbles. **Looney alert**
The genetic distance between Dresden/Munich and Lodz/Warsaw here is 0.00012. And these Poles aren't even from near Germany.
And yes, Germany has millions and millions of Germans of Slavic descent. Some ancient, of Wendish origin, and some recent, of Polish origin.
This German report proves it...
http://www.biotype.de/files/Immel_EJHG_06.pdf
But yes, the big difference between Poles and Germans is that Poles have a lot less affinity to Africa and Asia.
See Table 1. and Figure 2. in this current study.
It's something I've always suspected, and now its been proven by huge samples from both countries.
Ahh, I see...so you're Semitic?
Well in that case you can be thankful for the Polish and other Slavic input into Germany, as we not only Aryanized but also Europeanized your country.
And no, you can't twist history and reality. Slavic surnames reach 50% in parts of East Germany.
If you missed it...
http://www.biotype.de/files/Immel_EJHG_06.pdf
P.S. I'll be taking Dienekes to the cleaners here when new data comes out of Poland, on the relationship between Polish regions and surrounding nations. That stuff is on the way.
I think you'll enjoy it too. :0)
Yeah, Dresden Germans are just western Slovaks.
Difference in this study is 0.0005
Pft...
Poland is strongly Catholic even today and East Germany was completely Protestant.
But that cannot apply to the Middle Ages, when most of German colonization (and assimilation) happened. Furthermore, Poland-Lithuania was after that the most tolerant and multi-religious country of Europe. So you are still talking of mere recent events, not history.
Swabians have NEVER been Celtic, nor they are now.
Swabians as ethnicity are Germanic but their area was previously Celtic as most of Germany. Logically most natives were assimilated, maybe as serfs or slaves but assimilated anyhow.
In fact, the Franconian part of Bavaria is overwhelmingly Germanic while south of Danube shows more Celtic influence.
That may be due to Roman "pro-Celtic" intervention but still most Germany (excepted only Low Germany) has a Celtic background. Furthermore, Celts as ethnicity were formed there: somewhere in middle or upper Germany before they began invading the West.
Any pretension of total democide in the migrations of the past has absolutely nothing to be supported by.
'm not an expert but correct me if I'm wrong but:
http://dienekes.50webs.com/blog/archives/000050.html
http://dienekes.blogspot.com/2005/06/strong-differentiation-between-germans.html
Yes but most of those differences are due to Poland being more intensely influenced by Indoeuropeans early on (had much lower density in Neolithic "Danubian" times and was always more open to the East). The differences may have been emphasized by more "recent" historical trends such as the (west) German colonization fo the East or the even more recent ethnic cleansing of Germans in Poland and resettlement of Eastern Poles (from Belarus and Ukraine) in Silesia, Pommerania and other formerly German areas.
But still Germans are the main component of the bridge between Western and Eastern Europe as fig. 6 of this study clearly shows.
European population structure was carried out using high-density genetic variation on nearly 6000 individuals originating from across Europe. The individuals were collected as control samples and were genotyped with more than 3,00,000 SNPs in genome-wide association studies using the Illumina Infinium platform.
------------------
Sally
Transmitter
most Germany (excepted only Low Germany) has a Celtic background.
Ah, there it is again, that magical "Celtic" that means everything and nothing. Southern Germany was Celtic in a sense that it provided much of the cultural innovation at the time and/or was an active part of this pan-European movement, which was largely technological but also showed secondary administrative innovations. In Southern Germany, it had no linguistic nor genetic significance. Conversely, for many millennia before and after, there was a strong continuity in language and local genetic relationships, which is evident to this day.
These 2 peoples have always had a close relationship, though not always amicable.
Absolutely.
And for the gene freaks, northern Poles have a lot of influence from the most ancient Germanic people in the Baltic. Conversely, the "improper" (= not lower German, lower) Saxons, and also parts of Thuringia and Sachsen-Anhalt, have a deep history of being very pragmatic about the origin of their constituency over the past 3000+ years, or so.
At any rate, the Slavic expansion into this and surrounding regions was a very brief interlope of just about 300 years - that's it.
Before and after, dominantly Germanic tribes (which some Slavic influence) were the main, yet sparse settlers of this region --- a region that outside of some prime Loess localities was disliked since the beginning of agriculture for its harsh winters, drenching wet and cold springs, and poor drainage - to the point that it could not be properly worked with known species millennia after the western regions...
I can't believe what a few misguided words on a blog can achieve. Anyone still contemplating turning Swedes and Norwegains into seperate "races" please take into account the following...
If we're going to do a FINE SCALE genetic survey of Europe, then we also have to do some FINE SCALE sampling.
If we don't, then a lot of loonies will come out of the woodwork claiming Hitler had the foresight of 21st century genetecists, and so on.
It's funny, but kinda alarming at the same time.
I can't believe what a few misguided words on a blog can achieve. Anyone still contemplating turning Swedes and Norwegains into seperate "races" please take into account the following...
Swedes and Norwegians seem to be genetically separable. This was evidenced in the previous study (Lao) as well. Of course, a fine-scale mapping of Norwegian and Swedish populations may reveal overlaps, but that doesn't negate the distinctiveness of the two nations.
Ethnic groups have both a biological and a cultural component, so they are not simply "races". Biological distance implies some genetic isolation, and hence the necessary conditions for the emergence of cultural distinctiveness. Conversely cultural distance makes intermarriage more difficult and leads to biological distinctiveness.
Swedes and Norwegians aren't genetically seperable.
There's a huge blur between the two nations, and that's been missed due to poor sampling.
In fact, western Sweden has more in common with eastern Norway, than it does with eastern Sweden.
I repeat, this is fine scale genetics, and unless the sampling is as fine scale, then "racial" divides will appear where there are none.
Quothe Sound: "Very very few Poles can pass for Germans or vice versa."
Being American, and being exposed, as I have been, mostly to other Americans, I'm not in a position to dispute this, but I must say I find it hard to believe. What the heck is the difference in appearance between a German and a Pole? I'm part Polish, part German, part Swedish, part Irish, and part French-- and looking at my ancestors, the Swedes were the only ones which seem to have a phenotype that made them stand out from the others. (Being as they were huge and blond)
The others all looked more or less alike.
Swedes and Norwegians aren't genetically seperable.
There's a huge blur between the two nations, and that's been missed due to poor sampling.
Whether there is a huge or a small blur is an empirical question which will be determined by data. What we can say right now is that these results and those of Lao show a clear separation.
"Poland is strongly Catholic even today and East Germany was completely Protestant".
"cultural distance makes intermarriage more difficult and leads to biological distinctiveness".
So the difference between Slav and German is mainly a result of cultural factors.
Anyone here still supporting religion?
...the discovery that not only races, but even closely related ethnic groups (e.g. Norwegians and Swedes) can be distinguished with greater than 90% accuracy, serves to illustrate the scientific irrelevance of the ethnic nihilists and the affirmation that nations are, at least in part, genetic entities.
I would be hesitant to state that "ethnic nihilists" (or more accurately, race deniers) are scientifically irrelevant, since their ideas are still very prominent and taken seriously by mainstream scientists. It is racial realists, rather, in an era of racial hysteria, who have been made less relevant. What we can do is find/point to further evidence of the unscientific basis of the other side. What that can accomplish in practical terms is questionable.
"Mental hysteresis" [lagging?] lacks explanatory power. It is utopian ideals among White liberals along with attempts by cohesive minority groups to advance their ethnic genetic interests that are at the root of race denial. By diminishing or negating the concept of race these groups try to shape discussions of race and especially social policy.
At a time when European nations are being inundated with Third World immigrants and are experiencing birthrates that are below replacement, and when said minority groups are showing vitality and cohesion unseen in native Europeans (and particularly among the effete intellectuals on both sides of this debate), is there really much value in pointing the finger and telling these "ethnic nihilists" that their scientific model is inferior? It would seem to me that the best way to affirm race is to preserve race.
It seems to me that science can serve 2 distinct purposes: (1) to advance our knowledge about the world in which we live (the more popular view), or (2) to promote our perceived selfish interests, individual or group-wise, through memes (the preferred strategy of the race deniers).
The biological roots of both of these tendencies, though, are not dissimilar, and they need not be mutually exclusive.
Ah, there it is again, that magical "Celtic" that means everything and nothing. Southern Germany was Celtic in a sense that it provided much of the cultural innovation at the time and/or was an active part of this pan-European movement, which was largely technological but also showed secondary administrative innovations. In Southern Germany, it had no linguistic nor genetic significance. Conversely, for many millennia before and after, there was a strong continuity in language and local genetic relationships, which is evident to this day.
I don't realy understand what you're saying. The area of formation of the Celtic culture and language was with all likehood in mddle/southern Germany (and German-speaking areas of Switzerland and Austria, as well as he Czech Republic in general). Conversely the Germanic urheimat is furthern North.
Admittedly there's no absolute certainty on where exactly did Celts coalesced (though the Rhin area appears as the best candidate) but in general all that area mentioned before was a relatively homogeneous cultural area before and after IE expansion and it tended to converge culturally once and again. When we find the Celts more certainly located at protohistory, (excepting some earlier expansions like Iberia and southern France) they still appear centered in that area of Central Europe north of the Alps and south of the Germanics of Low Germany and Scandinavia. From there they extended (La Tène cuture) into most of France, Britain, Northern Italy, parts of the Balcans, etc. Since c. 200 BCE though they begin suffering the pressure of Germanic tribes, who manage to conquer one after another many Celtic "oppida" (fortified towns), disorganizing their economy. Against this Germanic push (and its consequences) would the Romans intervene since Caesar, delaying Germanic expansion in the Roman area of interest for many centuries.
The result is that you may certainly find some differences between one and the other side of the Roman limes in Germany but that does not mean that other more northernly areas were not Celtic before the Germanic push and have not kept some kind of Celtic background in one way or another. This doesn't mean that Celts were anything "magic" or whatever: like everybody else, they were in turn the product of their own background, both IE and Danubian, and even Magdalenian before them.
False. The "Turkification" of Anatolia happened in 12th century by Seljuk Empire.
How's that different from what I say? The Turkification was in any case mostly a process of absorption and assimilation of Anatolian natives who, at the time of Turkish conquest, spoke primarily Greek. The process anyhow continued along the Ottoman period, very specially as the Empire needed infantery and traders and other occupations that the Turkish aristocratic horsemen would not easily fulfill. In any instance converting to Islam (and therefore adopting Turkish as your language) had many advantages, so many obviously followed that path at one time or another.
And again, there have been a large scale immigration to ANatolia from the east according to Greek, Turkish, Armenian, Assyrian and Georgian historians.
"Large scale"? You just have to look at Anatolian Turkish genetics (even the Y-DNA, more susceptible to change via elite domination) to realize that Turks are not substantially immigrants, much less Turkic immigrants from the steppe (that part cannot account for more than a extremely low, neary irrelevant, percentage).
Anatolia has been densely populated since at least the Neolithic and the succesive invasor waves, while all leaving some small mark, have never been able to radically alter the genetic landscape. That applies to Turks as to Hittites, to Phrygians as to Greeks, etc.
False again, Nazi Germany only wanted to germanize the "desired" blooded people, not everyone. They thought that the desired people were ancient Germans lost in the Slavic world (may be referring to East Germanic tribes).
According to Generalplan Ost, 50% of Czechs, 25% of Belorussians, 35% of Ukrainians, 100% of Estonians, a sizable portion of Latvians
Not "false", as you admit that a good deal of Eastern Europeans, including noting less than 50% of Czechs (on paper) were considered by the Nazis to be "Germanic" (what for them basically meant "Nordic", no matter that most Germans are not actually Nordic at all).
True that Nazis had many weirdo ideas (most of them psychotic and criminal - and scientifically absurd) but if even such extreme racist psychopaths were willing to compromise with reality in their imperialist nightmare, guess that we an infer that the much more pragmatic (and not racist at all) Medieval and Modern Germans were much more open to assmilate the conquered peoples. And that is what history tells us: that many Slavs from Austria, Eastern Germany and what is now Poland, etc. were just absorbed as Germans after due transition. Of course, the opposite also surely happened in the previous stage of Slavic expansion (and so on till the Paleolithic).
Turks are a very heterogeneous people. It must be remembered that Turks are to a large extent an invented people, an attempt by the early 20th c. nascent Turkish nationalism to create a National identity out of its Muslim subjects.
Maju, do you even know what the Y-DNA pools of Central Asian populations are like? You appear to have no idea what you are talking about here.
Yes I do have an idea and in many cases they appear to be only partly Turkic/East Asian. The main exception would be Khazaks, where Eastern Y-DNA is clearly dominant. That's logical, because of the different enviroments and original settler density these different ethnicities/republics represent (for instance Uzbekistan has a very ancient presence, because of being an area much better suited for agriculture than the steppe).
You may know (not sure if you actually do) that the original homeland of Turkic peoples was roughly in modern Mongolia (and Mongols are from farther east in fact). The original Turks were East Asian though they obviously admixed intensely in the many centuries they dwelt in Central Asia and Eastern Europe. Whatever the case, I have absolutely no reason to look at the Anatolian, say, Y-DNA pool and think it's off its place in the West Asian/SE European context, not at all. But has almost no presence of typical East Asian halogroups such as C or O. Phenotypically, Turks also appear very much within their geographic context. You may easily confuse a Turk with a Greek, German or Iraqi... but hardly with a Khazak.
Sound of the Occident wrote: "You may be forgetting European history while you're living in Australia?"
From this part of the world (not actually Australia, but near enough) your arguments over differences within Europe do seem rather petty.
Now, Dienekes, and the rest of you Occidentals, hehe...
the point is, SNP clustering can't define races. At two clusters, humans fall into African-West Eurasian and an East Eurasian-American pools. This fine enough; we can say that these are two macro-races that can be broken down into smaller races. But the problem is there is no natural point where ever finer, more numerous clusters cease to become nonsensical, where we can define race. Clusters can go on into predicting ethnic groups, we we can see now, and the trend is that if you use even more SNP data, we can predict the region someone comes from, down to the village, the clan, for those places where there is still a lack of mobility.
Now, if clusters can separate two families, does that mean they are two different races?
There is no natural point, finer resolution where SNP clusters stop to make sense beyond the regional, ethnic level, as it can go on to cluster down to towns, villages, clans. This is why it can't define race and why Swedes and Norwegians are not sub-races.
Above is pure logic... but something tells me you guys are too busy to understand...
And some are just not intelligent enough... though it won't stop them from posting their 2 cents on here...
...if you use even more SNP data, we can predict the region someone comes from, down to the village, the clan...
Now, if clusters can separate two families, does that mean they are two different races?
Exactly what I said, while worded differently: "...basically distinguishing what is left over and commingled from extended families of the agricultural expansion. That's not races, that's cousins who outmarried."
In the context of:
"It's curious though that by Fst Rumanians appear pretty much distant from Poles and Russians (more than to Spain and UK) what makes them appear somewhat "western" in the whole context (also reflected in the graph)."
Maju said: Dacians did not spoke Gallic. The relation of Gaelic with Latin is not straightforward either. The reason why Romania speaks a romance language and not Slavic, Turkic or Hungarian beats me really. It's one of those unimaginable oddities of history, specially if you ponder that Dacia was part of the Roman Empire for a rather short period and had no Catholic influence...
...I have not the slightest idea why do you associate Romania with the Alps and Celts, really.
Long before Hun, Slav, Langobard, and Goth invasion, the vast area of the Illyrians was influenced by Celtic rulership and language in the north and northwest (Noricum), and by Roman intrusion and language from the west and south. Eventually, even Noricum was fully romanized. Here you have an example of a (at the time, just recently) Celtic speaking population, and one whose language is unknown but very likely was centum IE - both were romanized in short time likely due to the fact that their language had close affinity, in the first place - similar to the situation in cisalpine Gallia and in Rhaetia.
In fact, if you think about the origin of Latin: you had centum Celtic to the west and north, and centum Illyrian and Greek to the East. Makes sense that ~2,500 to 2,000 years ago, these were still fairly close to each other. I wouldn't be surprised if the southern pan-alpine Celtic at one point was almost as close to Latin as it was remote from Brythonic and island Celtic.
We don't currently know for sure the complex ethnic makeup of Romanians, and how much of a continuity from (satem IE) Dacians actually persists to the presence. But in addition to elements form the invaders listed above, and remaining groups as well as Latin-speaking Black Sea harbor inhabitants, it makes sense to me that there is also this large former Illyrian group: people who fled the above invaders by moving just a couple of hundred miles east, into the (after Roman retreat) sparsely populated and easily defensible deep forests and mountains of Romania. In fact, what other place did these people have, to go to? One thing we know for sure: they (conveniently) already spoke the vulgar Latin from which Romanian is derived, and for longer and more easily so than the Dacians.
I am sure we will know more in just a few years.
Eurologist: I can agree in principle with Latin/Italic being originally related to Celtic and Illyrian (though we just know too little of Illyrian and of Italic languages other than Latin) in the context of Central European cultural complexes (Tumuli-Urnfields-Hallstatt) pre-dating protohistorical Celts. But it's a debatable matter in any case.
But how do you relate those with Dacians, genenrally assumed to be closer to Thracians or to Eastern European groups, still beats me.
We don't currently know for sure the complex ethnic makeup of Romanians, and how much of a continuity from (satem IE) Dacians actually persists to the presence.
Probably all or most. Sure, they were once and again invaded by other peoples but none of them appears to have left much of a mark, excepting Romans (language) and Hungarians (plenty of minority settlements) maybe. In any case all those invaders, except Romans, have an Eastern origin. That cannot explain that they appear relatively "western".
Actually, from other studies Rumanians are not really distinct from other Balcanic peoples, so I think you should stop looking at them as "Latin" and begin looking at them as the only representative of the Balcans in this study.
...
At two clusters, humans fall into African-West Eurasian and an East Eurasian-American pools.
Hmmm, isn't it more like Tropical Africans and Eurasians+Oceanians+Americans? You never stop surprising me, Ren, sincerely.
Now I'd agree that the major divide within the Eurasian-plus pool is among Western (South Asians included) and Eastern (Oceanians and Native Americans included) sets. That's what the data I have seen once and again says.
Maju wrote,
"Hmmm, isn't it more like Tropical Africans and Eurasians+Oceanians+Americans? You never stop surprising me, Ren, sincerely."
The dividing line is indeed between Eura-Africans and Amerasians in the case of SNP clusters.
Grow up.
Grow up you man (you're incredibly arrogant and Sinocentric, you should be more balanced and, yes, mature).
There's no simple dividing line (there are many and many "bridges" too) but anyhow the main division fund in ALL studies is between Africans and non-Africans, what is in full ageement with everything we know. Now, I'd agree that I feel more identified instictively with Africans than East Asians often (probably because they have big round eyes, like most West Eurasians - but that's just a minor matter). I'd also agree that West Eurasians are somewhat closer to Tropical Africans than East Eruasians... but both Eurasian groups are overall closer to each other anyhow.
The dividing line is indeed between Eura-Africans and Amerasians in the case of SNP clusters.
That is correct, although it does not reflect the phylogeny in which Caucasoids and Mongoloids are more related to each other than they are to Negroids. The reason is quite simple: the HGDP panel is heavy on Caucasoid and Mongoloid populations and light on Sub-Saharans, so the first split capturing most information is between Caucasoids and Mongoloids; Negroids get bundled with Caucasoids since they are indeed closer to them than they are to Mongoloids.
Maju said,
"Yes I do have an idea and in many cases they appear to be only partly Turkic/East Asian. The main exception would be Khazaks, where Eastern Y-DNA is clearly dominant. That's logical, because of the different enviroments and original settler density these different ethnicities/republics represent (for instance Uzbekistan has a very ancient presence, because of being an area much better suited for agriculture than the steppe)."
It was a rhetorical question, maju. I obviously know much more about the Y-DNA pools of Turkic populations than you do.
The earliest historically recorded Turkic populations are from southern Siberia (especially the area around the Altai Mountains and the headwaters of the Yenisei River) and Central Asia (especially the area of modern Kyrgyzstan). Very few Turkic populations have ever dwelt in East Asia.
The haplogroup that is most widely and frequently shared among Turkic populations is R1a1-M17, with haplogroups J2 and R1b also being quite common among them. The majority of Kazakh males belong to haplogroup C3c, but this haplogroup is practically absent from most other Turkic populations, and it likely reflects a Mongolic, rather than Turkic, origin of Kazakh patrilineal ancestors.
If you find data on the Y-DNA of ancient Turkic populations that demonstrate conclusively that they possessed "East Asian haplogroups" rather than the haplogroups possessed by modern Turkic peoples, then I will give your claims some consideration.
Maju wrote,
"Grow up you man (you're incredibly arrogant and Sinocentric, you should be more balanced and, yes, mature)."
How many times have you attacked me with accusations before you ever found out the facts, calmed yourself down and thought about it?
Grow up. This message to get well shouldn't be coming from someone who is a lot younger than you.
Must be a specific paper I haven't checked myself (what I've seen does not seem to agree with that). Anyhow, what you say, Dienekes, is absolutely logical (it's an issue that pops up in all autosomal studies actually: sample size does matter a lot).
The former reply was meant for Deienekes. You guys post really fast.
To Ebizur: I'm following the quite mainstream thory that believes the Xiongnu as ancestors of the Turkic peoples, including the Huns. The Altai was, like Central Asia, prior to Hunnic/Turkic expansion an Indoeuropean region, so it's very hard to imagine the Turkic peoples pouring directly from there.
Very few Turkic populations have ever dwelt in East Asia.
Hmmm... Not just the Xiongnu but also the Göktürks controlled what is now Mongolia. They also expanded into Eastern Siberia (Yakuts).
The haplogroup that is most widely and frequently shared among Turkic populations is R1a1-M17, with haplogroups J2 and R1b also being quite common among them.
All those are Western Eurasian haplogroups clearly. R1a is widely accepted as an Indoeuropean marker, J2 as West Asian and R1b depends of subclade (the Central Asian subclade appears to have expanded with Tocharians, while the major subclade is clearly West Eurasian, mostly European).
What you are describing, in very misleading terms by the way (specially as Anatolian Turks make up almost 50% of all Turkic speakers worldwide), is the result of almost 2000 years of admixture with West Eurasian natives, who were obviously assimilated in most cases. The best "purebreed" representatives of the original Turks are surely Khazaks (if not Mongols, paradoxically), the rest are all (or most) basically absorbed or heavily admixed peoples.
he majority of Kazakh males belong to haplogroup C3c, but this haplogroup is practically absent from most other Turkic populations, and it likely reflects a Mongolic, rather than Turkic, origin of Kazakh patrilineal ancestors.
I'm not sure why Mongols and Turks (both micro-Altaic, a rather solid linguistic grouping) should be from different stock, moreso when the homeland of Turks appears to have been Mongolia. If anything it would show that Mongols have Turkic or Turkic-like patrilieal ancestors largely.
Also the Turkic expansion has taken many many centuries, while the Mongol expansion was a brief epysode, mostly manned by Turks.
If you find data on the Y-DNA of ancient Turkic populations that demonstrate conclusively that they possessed "East Asian haplogroups" rather than the haplogroups possessed by modern Turkic peoples, then I will give your claims some consideration.
If the Xiongnu are not Turks for you, then it will be difficult. I dont know of any such studies anyhow. You seem to want to limit the origin of Turkic peoples to just Göktürks, who were surely only a second moment in Turkic expansion (after Xiongnu/Huns) but I doubt that is valid at all.
Naturally the Turks that arrived at Anatolia were already mixed with West Eurasians but still the modern Turkish of West Asia (Azeris included) seem to be almost exclusively local Turkified peoples. After all, how would a buch of nomads alter meaningfully the genetic landscape of a densely populated agricultural area substantively? If they ever could, that would the exception, not the rule.
Some really smart comments here, particularly by those who further race-denial (a daring, iconoclastic position BTW) with the argument (forgive me if I get it wrong, I'm probably just not smart, patient, or whatever enough to understand, as suggested by "ren" several times above):
"The distinctions can be followed from the top (supposed "race") all the way down to the family level; are families races too?"
Now that is effing brilliant, not remarkably stupid or anything like that.
THAT's the way to stick it to racists; prove that race is analogous to family, with the difference being primarily one of scale.
Can you guys recommend some study guides, you know, where you learned your tricks of the trade...some source I can use to become just as clever? Other examples include ren's brilliant "I'd explain how your logic's circular, but you wouldn't understand," and "those scientists have poor reputations in the press," both instant classics!
Ren is right in this. I argue a lot with him but he's right in differences being gradual.
Additionally, I'd say that they are somewhat chaotic, as ancestry is very complex in every case, and genes recombine at every generation (not counting mutations).
Racialists, and specially these aggresive and sardonic racists, want humans to be like dogs, neatly divided in rottweilers, pekinese and terriers (among other groups) but forget that the very creation of these races is due to extreme and persistent practices of intentional breeding, exterminating all the "undesirable" offspring. This race breeding just does not exist among humans.
And anyhow, a dog is a dog is a dog.
We are just all "street dogs", or something very similar.
Additionally, I'd say that they are somewhat chaotic, as ancestry is very complex in every case, and genes recombine at every generation (not counting mutations).
Again, a devastating argument for race denial. Thanks!
Racialists, and specially these aggresive and sardonic racists, want humans to be like dogs, neatly divided in rottweilers, pekinese and terriers (among other groups)
Which is clearly ridiculous because Rottweilers, Pekinese, and terriers can't interbreed! And even if they could, Rottweilers, Pekinese, and terriers don't exist! It's all the same species, and it all comes down to clines and peaks and valleys and chaos and such. Any dog breeder can tell you as much. With a bit of training, a Poodle makes an excellent game dog.
but forget that the very creation of these races is due to extreme and persistent practices of intentional breeding, exterminating all the "undesirable" offspring. This race breeding just does not exist among humans.
Yeah, I mean everyone knows that humans don't self-select. I mean, even people in prison get conjugal visits.
And anyhow, a dog is a dog is a dog.
LOL! See? I just said the very same thing above! Great minds think alike!
Now, if I can only straighten out the artists. They think there's red, and orange, and blue, and stuff. Don't they know there's no clear, bright line between orange and red, and therefore there's no orange, and no red? In fact there's no way to distinguish where red ends and orange begins, but they go right on using these dangerous distinctions. Bunch of Philistines I tell you, but we'll get them thinking the right way eventually.
And the auto makers, now that I think about it. Who the hell do they think they are, with their "categories" of automobile? Cars, trucks, SUVs? What utter rot. What's an El Camino then? Pack of degenerates, but they'll get what's coming to them.
And since I'm on a roll, I think families should start looking over their shoulders, too. We're all mutts, street dogs, and a dog is a dog is a dog. Who is Mr. Jones to bequeath his worldly goods to his offspring? We're all the same, and Mr. Jones should share. There's no difference whatever between Mr. Jones' "children" and a Bantu. Jones has no right to discriminate in that way. Families! Bunch of Amalekites, I tell you. They'd better watch their step, we're going to have to straighten them out soon.
We are just all "street dogs", or something very similar.
Indeed. I see no differences between Bantus and Ashkenazi Jews. In fact, I think Israel should be filled with Bantus, right up to the brim.
Some might say that, if we're all the same, then what's the point of diversity, affirmative action, and all the rest, but I'm not one of those. Which reminds me, how come the American federal government is so racist as to categorize people into "black," "white," "hispanic," etc., for the purposes of handing out favors? I mean, we're obviously all indistinguishable, so where do they get off? They're going to need sorting out as well, I suppose.
Sorry, I managed to accidentally edit out part of my response:
Ren is right in this. I argue a lot with him but he's right in differences being gradual.
He's right, and you've clearly put your finger on where I disagreed with him (obviously it wasn't some different point, about how the distinctions involving race scale up and down from the macro to the micro, and how that makes race and family analogous, and does nothing whatever for race denial, or anything like that), so my mistake. I'm trying to learn from you guys, but I have my work cut out for me, that's for sure!
Btw, I think it's great that all these studies come with graphs, plotting the Xs and Ys of where "members" of "groups" fall.
Those clusters of "members" of "groups," each with their own color, each falling in their own spaces on the graphs, REALLY DRIVES HOME how we're all the same.
Cheers!
Which is clearly ridiculous because Rottweilers, Pekinese, and terriers can't interbreed!
What really shows your ignorance only. All dogs can interbreed (though guess there can be some difficulties if there's important size difference) and they actually do. They can also interbreed with wolves and even with foxes (though in the last case at least the hybrids are definitively not domestic: they will eat your chicken with all likehood).
I see no differences between Bantus and Ashkenazi Jews.
Certainly not many. I come from a family where, as in many other European families, you can find almost all shades of skin color, presence and absence of epicanthic fold, nearly all kind of hair color and textures, not to mention eyes: from deep brown to light blue. And we all share a good deal of the tiny variable fraction of human genes (roughly 50% with 1st degree relatives, 25% with 2nd degree, etc., though there is randomness going on).
So what does make me similar to any other European, Basque or whatever? They are not my direct or even distant relatives in most cases, our genetic share is surely virtually null beyond what makes us humans. And yes that's like 99% (can't recall) - but that's precisely the non-variable part of the genome.
There are a handful of genes that because founder effects, drift or whatever have become regionally fixated or common, while elsewhere they are rare. Can you tell me, Svastikos, what percentage of the human genome do they make? It must be a extremely low ammount, probably less of what separates me and my mother (who according to your shallow visual perception should be of different races, no matter that we share some 50% of all variable genes).
Which reminds me, how come the American federal government is so racist as to categorize people into "black," "white," "hispanic," etc.
I agree that's racist, and the British do it too, btw, probably because of US influence - I read that in the past "race" in Britain, at least in the army, meant: Eglish, Welsh, Scottish or Irish. "Jewish? That race does not exist, boy. That's a religion".
Those clusters of "members" of "groups," each with their own color, each falling in their own spaces on the graphs, REALLY DRIVES HOME how we're all the same.
Those graphics account only for a minimal ammount of the whole genome. The very vast majority is shared, so it gves no info about the differences and is discarded. All variability that does not cluster is also discarded (PCs seldom represent more than 10 or 20% of all the variable genes - never did my homewrok for this but it's like that). The result, representing the minimal regionally clustered variability is what is plotted. Nothing else.
You can get even lower level plots, you can get clusters even in such a homogeneous region as Europe or among Native Americans. And guess you would get clusters in even tiny homogeneous territories like Iceland or Luxemburg. You get them for Finland and Scandinavia certainly.
There's always some alelles that are more common here than there. They would appear to show some founder effects of sorts or homogeneization along time (drift) but they always represent only a tiny part of the human genome necesarily.
If you get Japanese, Nigerians, Britons and chimpanzees you will necesarily see that all humans cluster at one extreme of the PC1, and very tightly. In fact, that's what happened when they compared Neanderthal and H. sapiens' mtDNA: the main difference, by a lot, was between Neanders and all humans, living or fossil.
Clusters and PCs are relative. You need a perspective to know what they really mean.
But if you have an ideology, you don't care: you'll look for anything that justifies it. When you have an ideology like yours the goal justifies any methods, but when you approach things with scientific mind it's the methods what matter, as there is no goal but the truth.
What really shows your ignorance only. All dogs can interbreed (though guess there can be some difficulties if there's important size difference) and they actually do.
They can? What do they call these interbreed mixes? I never heard of such a thing! Gosh, I guess that shows my ignorance plainly! I sure have a lot to learn!
Guess that proves how different dog breeds and human races are for me, right there!
I see no differences between Bantus and Ashkenazi Jews.
Certainly not many. I come from a family where, as in many other European families, you can find almost all shades of skin color, presence and absence of epicanthic fold, nearly all kind of hair color and textures, not to mention eyes: from deep brown to light blue. And we all share a good deal of the tiny variable fraction of human genes (roughly 50% with 1st degree relatives, 25% with 2nd degree, etc., though there is randomness going on).
See, this is the kind of rigorous scientific thinking I come to Dienekes blog for! "My family's mixed, ergo, Bantus and Ashkenazis are the same"! Profound!
So what does make me similar to any other European, Basque or whatever? They are not my direct or even distant relatives in most cases, our genetic share is surely virtually null beyond what makes us humans. And yes that's like 99% (can't recall) - but that's precisely the non-variable part of the genome.
Brilliant! If you have no race, I have no race! I'm beginning to understand now.
Numerically, -100,000,000 and 100,000,000 are 90% the same! One little symbol in the front is changed, and that's it! Ergo, it makes no difference which figure represents your ledger for the past year! Brilliant! Profound! It just keeps getting better and better with you guys!
Maju, do you mind if I quote you on other sites from time to time?
There are a handful of genes that because founder effects, drift or whatever have become regionally fixated or common, while elsewhere they are rare. Can you tell me, Svastikos, what percentage of the human genome do they make? It must be a extremely low ammount, probably less of what separates me and my mother (who according to your shallow visual perception should be of different races, no matter that we share some 50% of all variable genes).
Yeah, Svastikos should answer your penetrating questions immediately. I want to know how that bastard responds. I mean clearly, if we can show that the genes that make Rottweilers Rottweilers and Poodles Poodles are some ostensibly small percentage of the total genetic makeup of Rottweilers and Poodles, then obviously dog breeds (and races!) don't exist.
This should be obvious! I'm not nearly as brilliant and erudite as Maju, but I think I can help against this racist Nazi bastard Svastikos:
Humans And Chimps Differ At Level Of Gene Splicing
Researchers are closer to understanding why humans differ so greatly from chimpanzees in the way they look, behave, think, and fight off disease, despite having genes that are nearly 99% identical.
Never mind all that mumbo-jumbo about expressed differences, the point is that Humans and Chimps are 99% the same.
Most human-chimp differences due to gene regulation--not genes
The vast differences between humans and chimpanzees are due more to changes in gene regulation than differences in individual genes themselves...That 1975 paper documented the 99-percent similarity of genes from humans and chimps and suggested that altered gene regulation, rather than changes in coding, might explain how so few genetic changes could produce the wide anatomic and behavioral differences between the two.
Never mind any of that crap about "vast differences" and "wide anatomic and behavioral differences" between Humans and Chimps. The point is, Humans and Chimps are 99% similar.
So there you have it. Humans and chimps do not exist! So OBVIOUSLY human races (and "population groups," for you lab-coat-wearing racists) don't exist (this is the kind of logic I'm learning here, so I want to repeat how grateful I am for the help of honest brokers of information and science, like Maju and Ren).
Which reminds me, how come the American federal government is so racist as to categorize people into "black," "white," "hispanic," etc.
I agree that's racist, and the British do it too, btw, probably because of US influence - I read that in the past "race" in Britain, at least in the army, meant: Eglish, Welsh, Scottish or Irish. "Jewish? That race does not exist, boy. That's a religion".
Of course we agree. This only serves to highlight the good sense of our open, honest, well-intentioned (I mean, I certainly know better than to think you've assayed your family, and projected the results onto the world in an attempt at self-legitimization) discussion, while the world organizes around something that doesn't exist. I think it's almost as significant as Nero fiddling while Rome burned.
Those clusters of "members" of "groups," each with their own color, each falling in their own spaces on the graphs, REALLY DRIVES HOME how we're all the same.
Those graphics account only for a minimal ammount of the whole genome.
Indeed! Which means that race does not exist, obviously, as I showed above with my brilliant Human/Chimp analogy.
But if you have an ideology, you don't care: you'll look for anything that justifies it. When you have an ideology like yours the goal justifies any methods, but when you approach things with scientific mind it's the methods what matter, as there is no goal but the truth.
I agree. When you have this ideology blinding you, you "see" race. When you don't, you can see clearly enough to NOT project assays of your family onto the world. You can see that Human and Chimp are the same!
Clearly, all the honest brokers are on our side, because we're all about science. And what a cozy coincidence that our position is that of the western zeitgeist and elite. How often does your teevee tell you EXACTLY what the science tells you (AND was telling you for decades before genetics had anything much to say)? And just as clearly, all the dishonest, ideologically blinded sorts fall on the other side of the argument (maybe that's why most sites censor this kind of argument? Because they know better than to waste time with these sorts?). Those bastards!
Oh, Maju - I try to do my own homework, but I confess I'm not as bright as you. Could you tell me what this Svastikos' ideology is? I mean, I don't have a clue beyond following what I perceived to be your lead (e.g., he's a racist Nazi bastard), but I'd like to have more than that if he shows up again. What IS his ideology, precisely? I ask because you seem to have a pretty good handle on it; you know enough about it to know this racist Nazi bastard is blinded and stuff, so I figure you have a flying, cartwheeling, backflipping clue as to what that ideology is. Could you elaborate?
Btw Maju, I know it must be my fault because you're so scientifically-minded, and obviously smarter and more learned than I, but you didn't respond to my idea of flooding Israel with Bantus. Personally I think it's a marvelous idea, but I'd like your feedback.
They can?
Yes, they can.
What do they call these interbreed mixes?
Dogs.
My family's mixed, ergo...
All families are mixed. There's no such thing as natural clonation in humans (except for identical twins).
If you have no race, I have no race!
You have no race: it's just an idea in your mind. Don't be ridiculous.
Numerically, -100,000,000 and 100,000,000 are 90% the same!
I'm not saying that. And if you knew anything about genetics you'd know.
If humans would be 90% different, we would not be the same species. Such extreme differences can maybe exist between different biological kingdoms like plants and animals. Only that. Even chimps are more than 95% like us humans (from memry something like 98.5% actually). That means that all humans are at least 99.5% the same, probably more like 99.9%.
You are counting the difference between 1 billion sand grains and 1 billion and one. There's one grain of difference but only an obsessed mind can give it any importance at all.
Maju, do you mind if...
Well, to be honest, I mind that people like you exists alltogether but can't do much about it, except trying keep logic on, to prevent the cancer of irrational ideologies from spreading and going back to the times of the Inquisition and the Holocaust.
Sincerely I have no hope of persuading you: you do not think rationally, just use pseudo-logic to justify an ideology. If the world would work like people like you claim, Achilles would never reach the turtle.
if we can show that the genes that make Rottweilers Rottweilers and Poodles Poodles are some ostensibly small percentage of the total genetic makeup of Rottweilers and Poodles, then obviously dog breeds (and races!) don't exist.
Do they? Do dogs care about that? Obviously not (except maybe in matters of size, what in their world may mean power). Humans do not show such extreme size differences and even the extremes are not even twice the size of each other. This shows how dog races are a product of extreme artificial selection and only that and that, would dogs be allowed to reproduce freely, as humans mostly do, such races would quickly disappear.
This should be obvious! I'm not nearly as brilliant and erudite as Maju, but I think I can help against this racist Nazi bastard Svastikos:
Humans And Chimps Differ At Level Of Gene Splicing
Interesting but... so what? I tend to agree that a lot behind phenotypes is epigenetic and not merely genetic but this field is still in its infancy. We can't judge what we still know so little about (and I remind you that most of epigenetic effects are enviromentally influenced).
The more important epigenetics is, the less relevant genetically defined "races" would be.
Never mind all that mumbo-jumbo about expressed differences, the point is that Humans and Chimps are 99% the same.
Almost. More like 98.5%. This is no surprise, right? We share so many things with human and bonobos that no other animal has... I don't care if they are low vaulted an hairy, they are so terribly human anyhow!
Never mind any of that crap about "vast differences" and "wide anatomic and behavioral differences" between Humans and Chimps. The point is, Humans and Chimps are 99% similar.
Actually, if you think about it, we are also extremely similar in phenotype and behaviour. Maybe you like more lions or eagles for heraldic/ideal reasons... but these animals (still relatively close to us in comparison to, say, grasshoppers or sharks) are "ETs" when compared to the Pan/Homo genus.
It seems to me that you're being extremely homocentric here, probably bacause of the Judaistic background of modern European culture and all that legend of being made "to the likehood of god". Such detachment from Nature is sick and has many negative consequences, beginning with a bad understanding of who we are, where we come from and what the heck are we doing here in life.
So there you have it. Humans and chimps do not exist!
We have been separated by like 8 million years, possibly more. In comparison humankind has only existed for some 200,000 years and the groups you now call "races" only began to diverge some 120-60,000 years ago (depending on the model). So, in terms of time, differences between humans are of four or rather five orders of magnitude the differences between chimps/bonobos and us.
So, yes, if we take humankind as reference (and not the whole animal kingdom or even mammals or even primates... or even great apes), humans an chimps/bonobos are pretty much different.
But you cannot use those differences to argue for the ones we find within humans. They are totally different scales. Galaxies and atoms again.
I certainly know better than to think you've assayed your family, and projected the results onto the world in an attempt at self-legitimization
I need to legitimaze nothing. Anyhow my full family would be considered "caucasian" (ahem: Caucasian are people from the Caucasus region, what you mean is "Caucasoid") by your racist standards. I also see that in so many other people all through Europe. I.e. documentary on Norwegian fishermen and the guy looks a tall Greek, so many Brits that look Spaniard, Scots that look Basque, Germans that look Italian... it's all around. Dunno, some Black Africans look like dark Caucasoids, while some Caucasoids look depygmented versions of Mongoloids or Polynesians, etc.
Your neatly packed boxes are nowhere: diversity and remote likeness is what I see more. I can see the differences you see too but they are not, never, in neatly packed boxes. Genes are just too wild for that, and phenotypes even wilder.
Indeed! Which means that race does not exist, obviously, as I showed above with my brilliant Human/Chimp analogy.
As mentioned it depends on the viewpoint. From a wide biological perspective chimps and humans are almost the same but from a narrow homocentric perspective, they are very different. But you have to be extremely race-centric to emphasize the ridiculous microscopic differences within humankind.
When compared with our closest known relatives, Neanderthals, humans (fossil and living) display differences that are very small in comparison. And Neanderthals were very much like ourselves. Even the most extreme human differential type (probably Mungo Man) was really close to the main human cluster in comparison, not with chimps, but with Neanders.
You just like to emphasize differences that are basically subjective. I bet you can stay hours arguing wether turqoise color is blue or green.
You can see that Human and Chimp are the same!
Almost the same.
How often does your teevee tell you...
I watch very little TV, honestly. It's boring.
Could you tell me what this Svastikos' ideology is?
Sure, I could...
Could you elaborate?
Maybe when you stop using sarcasm and allow normal discussion to happen.
... but you didn't respond to my idea of flooding Israel with Bantus
If you read my blog (what I'm sure you don't), you'd know that I'm against the very existence of Israel, which I think it's comparable to Nazi Germany or, maybe more properly, Apartheid South Africa. The last European (or should I say US-American?) colonial project, a neo-crusade, a doomed yet painfully racist project.
I am also against some Judaistic influences, specially Christian and Islamic religions. But I'm not anti-Semitic, as I admire many Jews, like Spinoza, Marx, Reich, Einstein, Chaplin... you know.
But Zionism is a crime against Humanity and the JCM religious complex that has dominated Western Eurasian culture/s in the last 2000 years is an insult to intelligence and a burden of sexual and intellectual repression that only gradually we manage to overcome now. It must be said that, among others, many Jews have helped in this path to Illustration and Humanism.
I'm balanced in all this: you can't really accuse me of antisemitism or semitism. I don't judge people because of their ethnicity, though I know that cultural background can be very influential.
this one is interesting! i really wonder where southern italians, sardinians and greeks woud be placed here on this map? more south and east then romanians? or more near iberians? hmm
Having been a part of the Online Universal Work Marketing team for 4 months now, I’m thankful for my fellow team members who have patiently shown me the ropes along the way and made me feel welcome
www.onlineuniversalwork.com
"Nonetheless, at a time when -due to a sort of mental hysteresis- proclamations that "races are social constructs" are still routinely made, the discovery that not only races, but even closely related ethnic groups (e.g. Norwegians and Swedes) can be distinguished with greater than 90% accuracy, serves to illustrate the scientific irrelevance of the ethnic nihilists and the affirmation that nations are, at least in part, genetic entities."
Sounds somewhat sensationalist and a bit of a strawman to me.
I think that the main point of the people who say that "races are social constructs" is that, unlike many people seem to think, the biological differences between populations aren't expected to explain every difference we see between racial "lines" (or county lines), not that no one would ever find genetic markers on populations that have split some time ago, and things like that. Instead, things like that can be as easy to accept as different fingerprints for different persons. It's almost like "populational fingerprints", useful as such, but as fingerprints, not useful to explain much more, except in the heads of believers of palmistry. Similarly, 99,999% of people who mention even gender as a social construct are probably not saying that men can get pregnant as well, but only that the common roles of each gender aren't as biologically hardwired as the barking in the dog and the meowing in the cat.
Post a Comment