UPDATE (Jan 10):
While previous work on the Y chromosomes of Russians had established the main conclusions (dominance of Y-haplogroup R1a, and a Finno-Ugrian N3 substratum), this paper adds to our understanding by examinining several ethnic Russian populations and placing their variation within the larger Eurasian context.
An interesting multi-dimensional scaling plot from the paper. For each ethnic group, the large disk indicates the entire group, while the smaller figures, geographical subpopulations. This is an interesting way to present the information, and shows clearly (a) the tight clustering of Slavic populations in a large area from Poland to Russia and the Ukraine, and also the evidence of the Russification of indigenous Finno-Ugrians (populations 1-4: Mezen, Pinega, Krasnoborsk, Vologda).
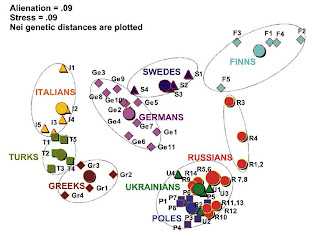
Also of interest for students of Slavic origins, another recent article about which I had blogged here. Note also the distance between all Greek subpopulations, including Macedonian Greeks from the Slavic cluster which should be read as further evidence contra the Fallmerayer thesis.
The Greek, Turkish, and Italian populations are well separated from the northern and eastern European populations on the left side of the figure; Germans are intermediate between southern Europeans and Swedes who tend to the Finns; like the northern Russians, Swedes also have their own Finnish influence. Evident, also, is the differentiation between Slavs and Germans, which had been noted before.
Also of interest from the paper is the comparison of inter-ethnic variation within European ethnic groups (also evident in the Figure):
Table 3 summarizes data on Y chromosomal intraethnic variation among Russians and compares them with other ethnicities of Europe. The highest variation among subpopulations is found for Finns, Croatians, Russians, and Italians (GST value between 0.04 and 0.08); Swedes and Germans demonstrate moderate variation; other ethnic groups (Greeks, Turks, Poles, Belorussians, and Ukrainians) exhibit similar and lower level of regional variation (GST value approximately 0.01).
Am J Hum Genet. 2008 Jan;82(1):236-50.
Two sources of the Russian patrilineal heritage in their eurasian context.
Balanovsky O, Rootsi S, Pshenichnov A, Kivisild T, Churnosov M, Evseeva I, Pocheshkhova E, Boldyreva M, Yankovsky N, Balanovska E, Villems R.
Progress in the mapping of population genetic substructure provides a core source of data for the reconstruction of the demographic history of our species and for the discovery of common signals relevant to disease research: These two aspects of enquiry overlap in their empirical data content and are especially informative at continental and subcontinental levels. In the present study of the variation of the Y chromosome pool of ethnic Russians, we show that the patrilineages within the pre-Ivan the Terrible historic borders of Russia have two main distinct sources. One of these antedates the linguistic split between West and East Slavonic-speaking people and is common for the two groups; the other is genetically highlighted by the pre-eminence of haplogroup (hg) N3 and is most parsimoniously explained by extensive assimilation of (or language change in) northeastern indigenous Finno-Ugric tribes. Although hg N3 is common for both East European and Siberian Y chromosomes, other typically Siberian or Mongolian hgs (Q and C) have negligible influence within the studied Russian Y chromosome pool. The distribution of all frequent Y chromosome haplogroups (which account for 95% of the Y chromosomal spectrum in Russians) follows a similar north-south clinal pattern among autosomal markers, apparent from synthetic maps. Multidimensional scaling (MDS) plots comparing intra ethnic and interethnic variation of Y chromosome in Europe show that although well detectable, intraethnic variation signals do not cross interethnic borders, except between Poles, Ukrainians, and central-southern Russians, thereby revealing their overwhelmingly shared patrilineal ancestry.
Link

1 comment:
what for BS its no assimilation but mixing. Its Finnish myth to get russian land. You can see that north russians are clearly distant like a mix. Russians have also their own haplogroup which comes from siebrian people while fins have one coming from far east.
http://i1311.photobucket.com/albums/s666/malaria2000/ejhg2008101f1_zpsf3ed01b5.jpg
Post a Comment